Metagenomics 16S analysis | BatchX - Supercharge your research with our end-to-end bioinformatics platform.

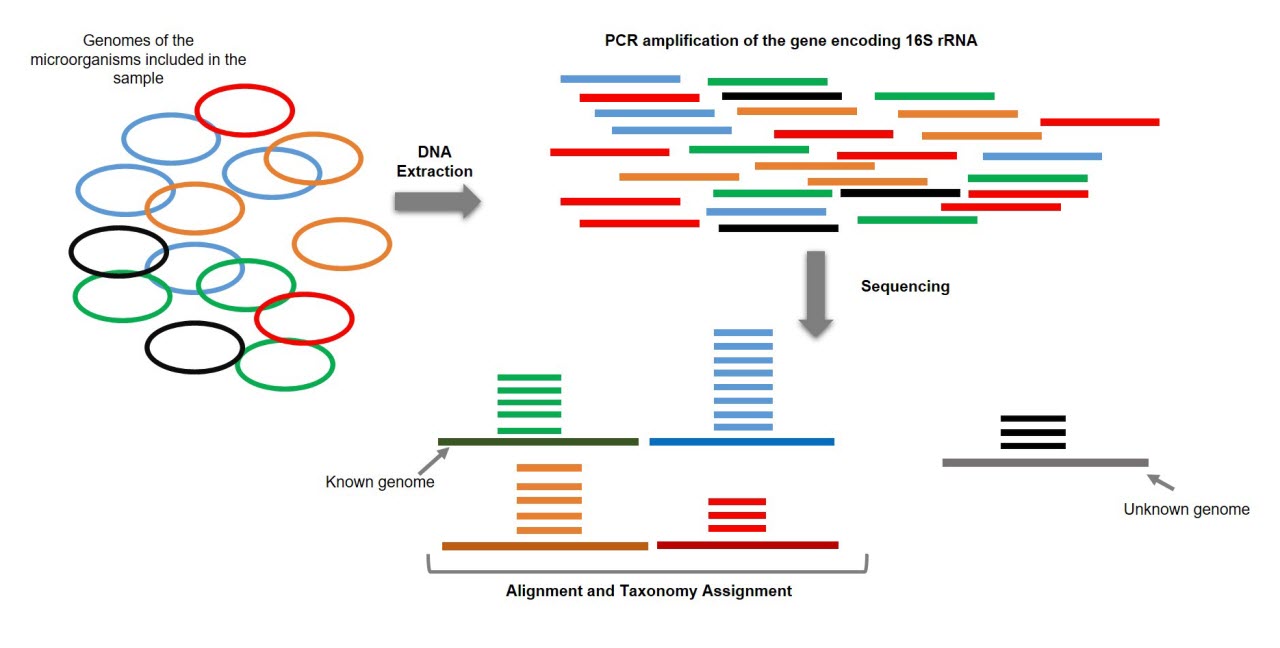
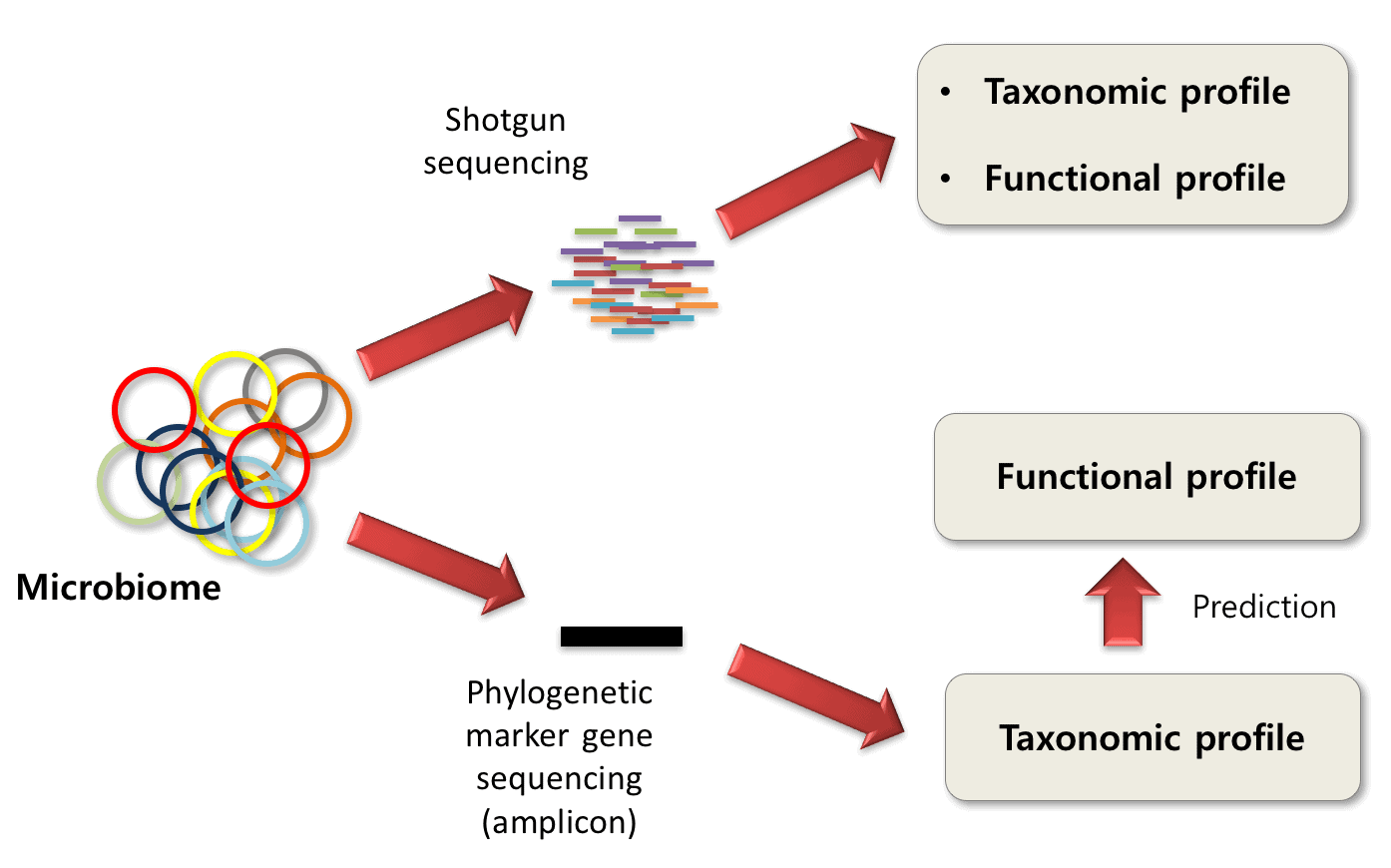
Estimating taxonomic profiles using 16S rRNA targeted sequencing or... | Download Scientific Diagram
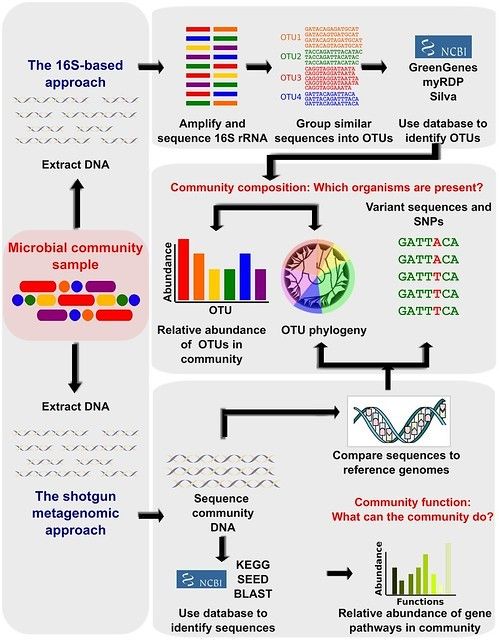
Blogpost shared by Microbiome Insights: "16S rRNA Gene Sequencing vs. Shotgun Metagenomic Sequencing"

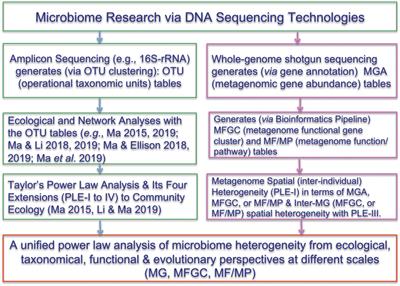
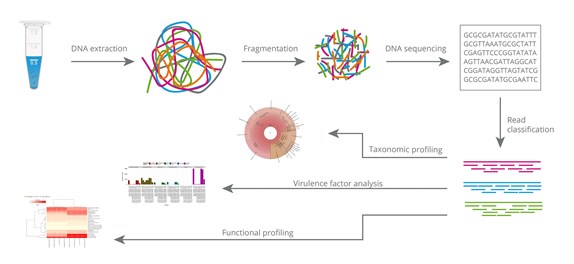
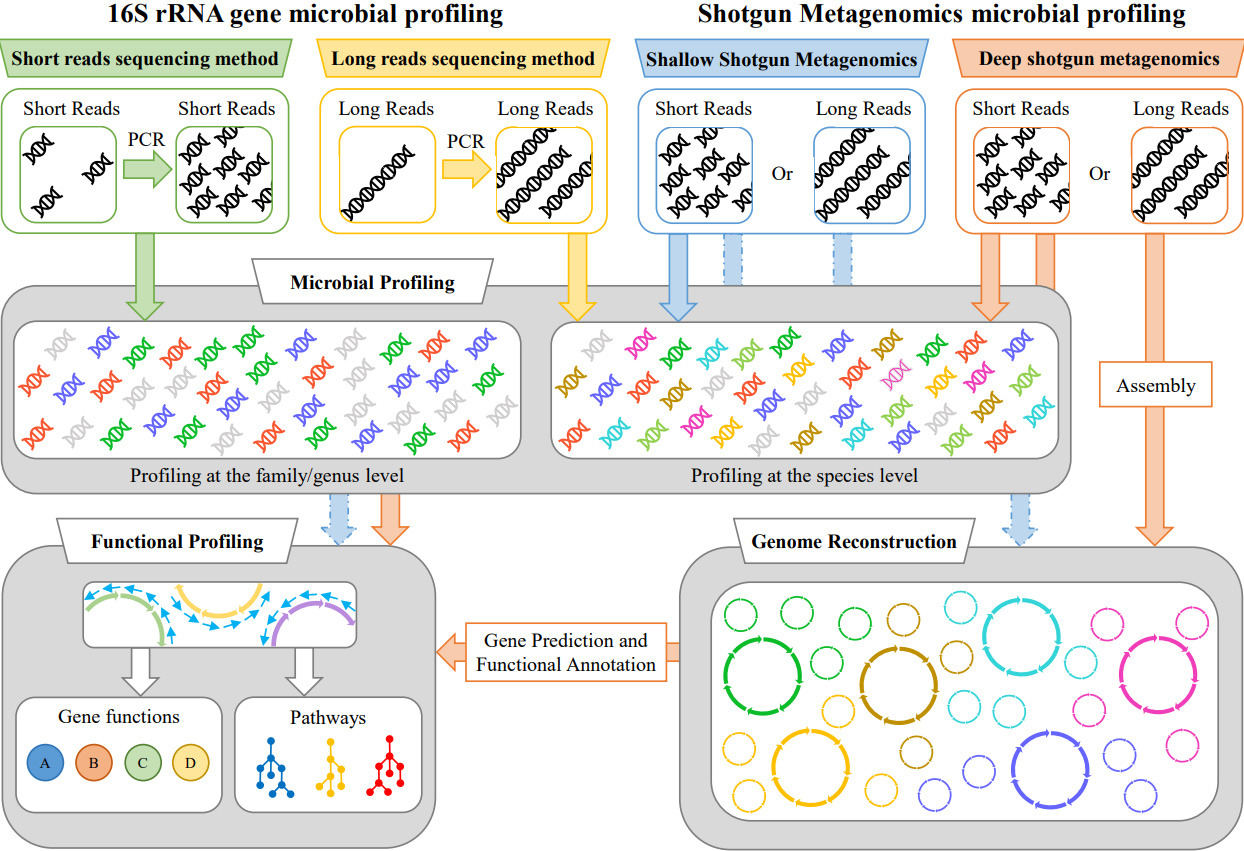
General overview of the bioinformatic pipelines for the 16S rRNA gene microbial profiling and shotgun metagenomics. General overview of the bioinformatic. - ppt download
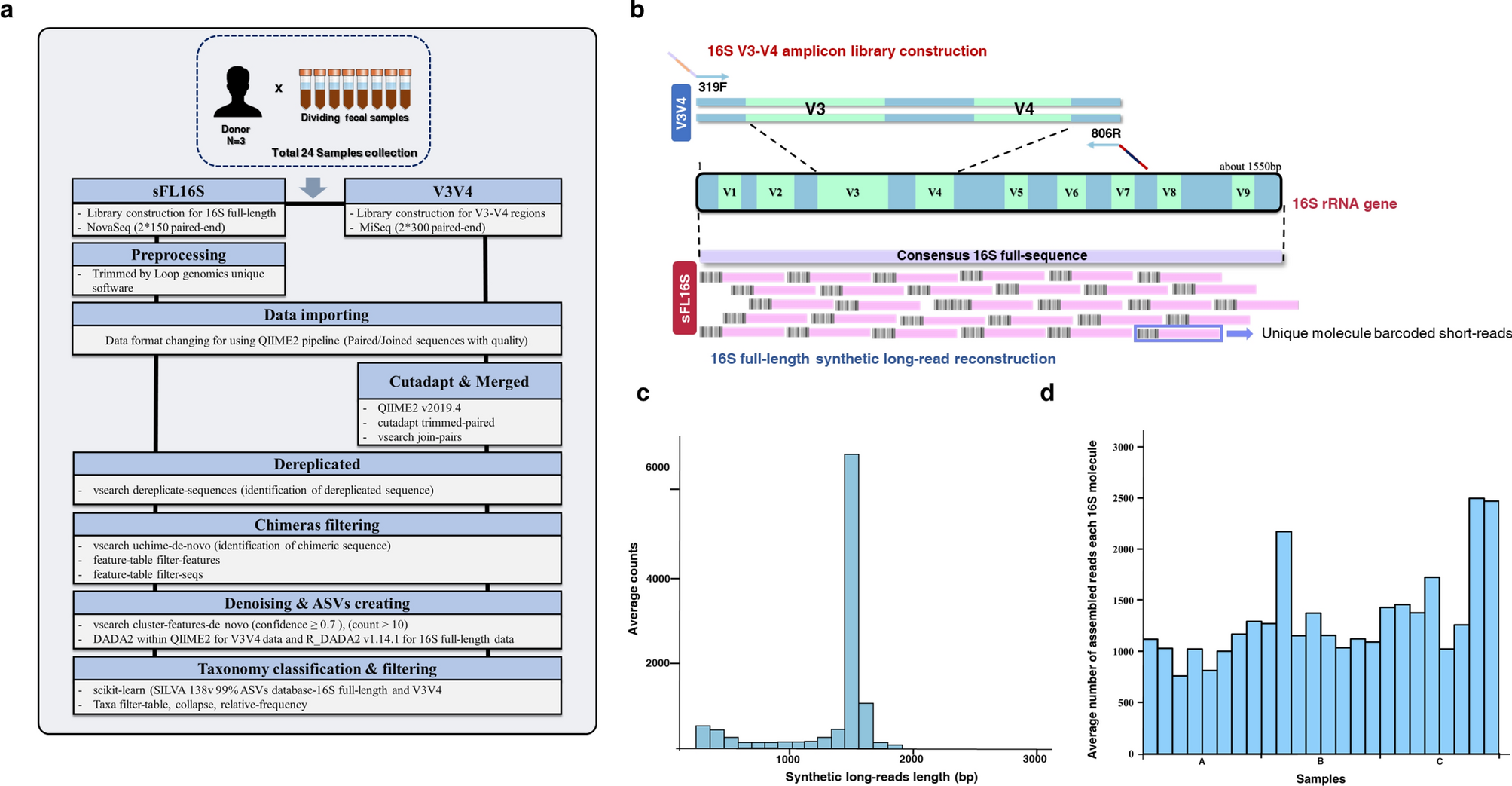
The effect of taxonomic classification by full-length 16S rRNA sequencing with a synthetic long-read technology | Scientific Reports

Comparison of reduced metagenome and 16S rRNA gene sequencing for determination of genetic diversity and mother-child overlap of the gut associated microbiota - ScienceDirect
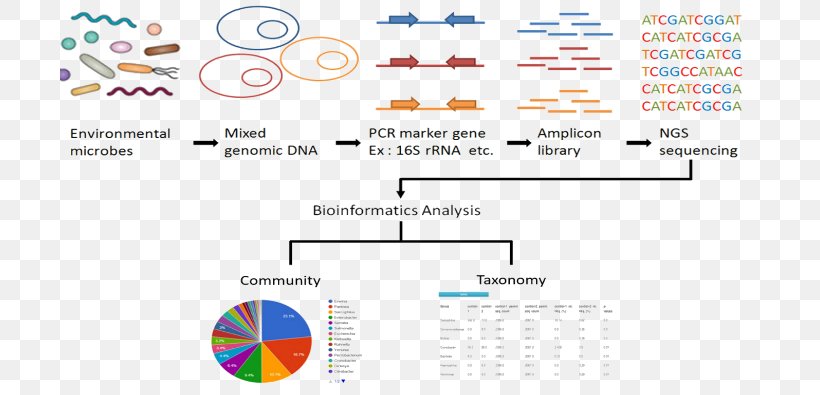
16S Ribosomal RNA Metagenomics Sequencing, PNG, 700x395px, Metagenomics, Amplicon, Archaeans, Area, Bacteria Download Free

Comprehensive 16S rRNA and metagenomic data from the gut microbiome of aging and rejuvenation mouse models | Scientific Data
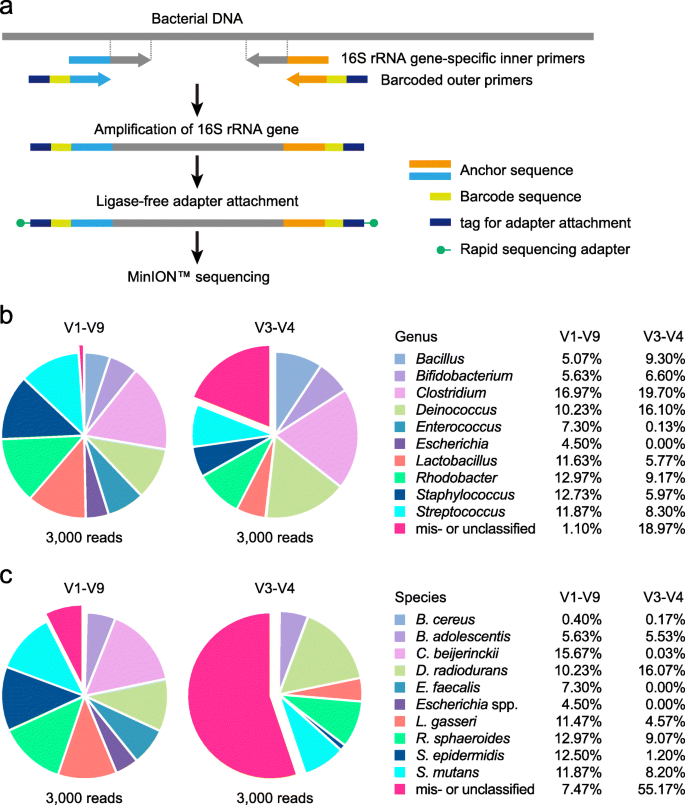
Full-length 16S rRNA gene amplicon analysis of human gut microbiota using MinION™ nanopore sequencing confers species-level resolution | BMC Microbiology | Full Text

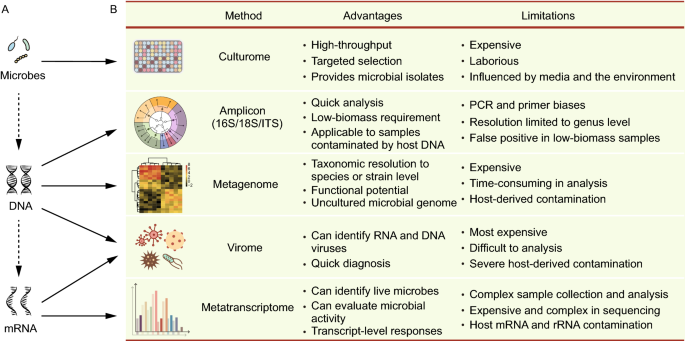
16S vs Shotgun Metagenomic Sequencing: Pros and Cons for Microbiome Studies | How do you choose between 16S rRNA gene sequencing and shotgun metagenomic sequencing for microbiome studies? There are many factors
A breath of fresh air in microbiome science: shallow shotgun metagenomics for a reliable disentangling of microbial ecosystems