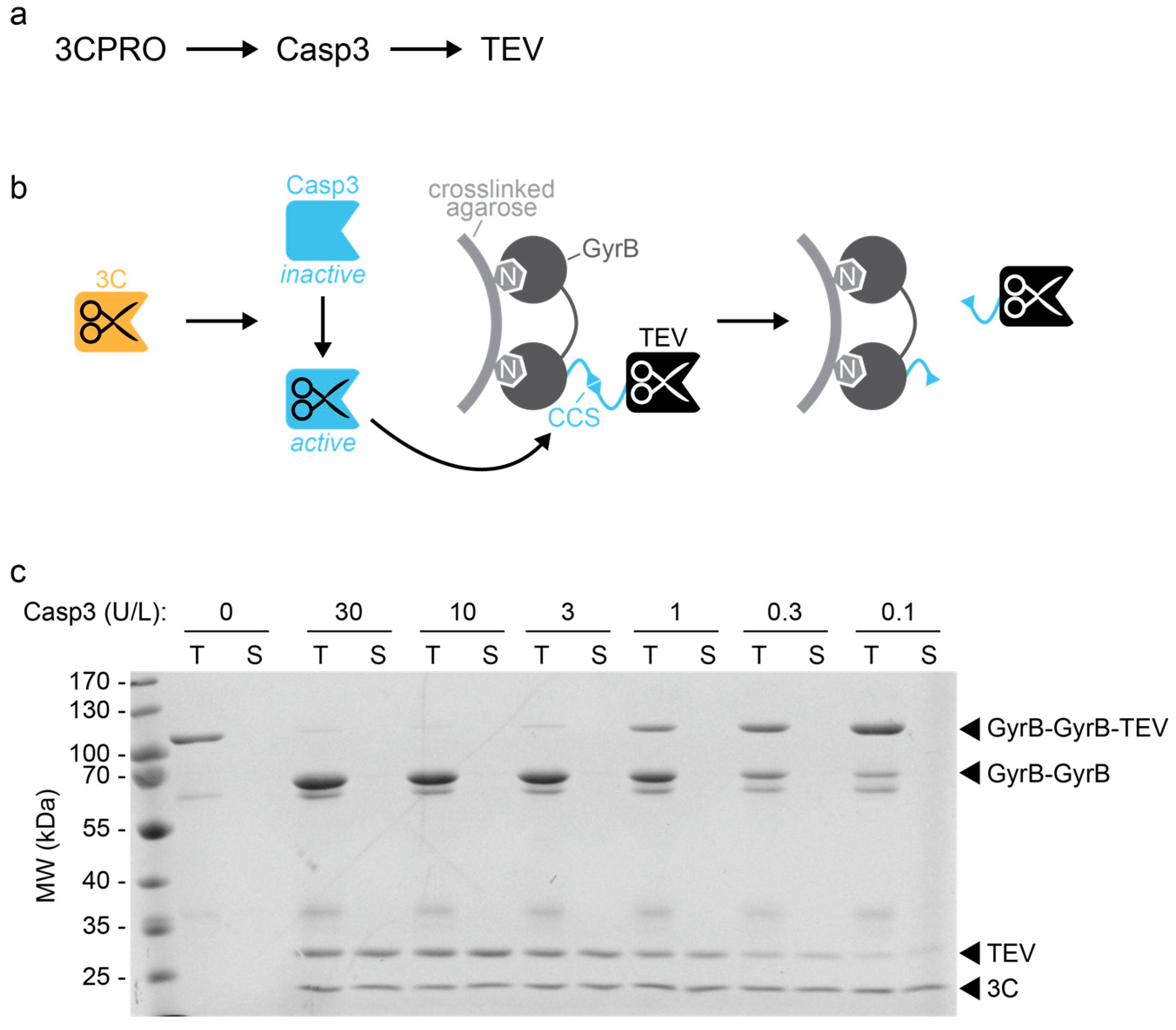
A split protease-E. coli ClpXP system quantifies protein–protein interactions in Escherichia coli cells | Communications Biology

A) Three tags, i.e. 10x-His, FLAG and HA, and one proteolytic site for... | Download Scientific Diagram

Human Rhinovirus 3C protease cleaves RIPK1, concurrent with caspase 8 activation | Scientific Reports

SDS-PAGE showing the HRV 3C protease activity in the presence various... | Download Scientific Diagram

SARS-CoV 3CLpro cleavage sites and the canonical recognition sequence.... | Download Scientific Diagram
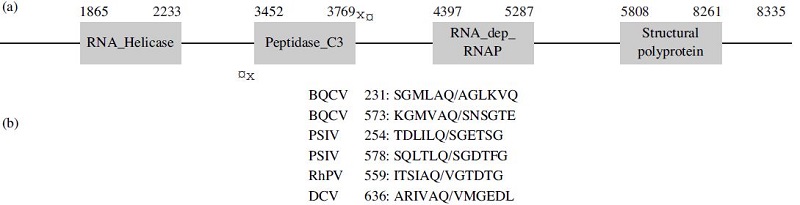
Frontiers | Identification of Cleavage Sites Recognized by the 3C-Like Cysteine Protease within the Two Polyproteins of Strawberry Mottle Virus
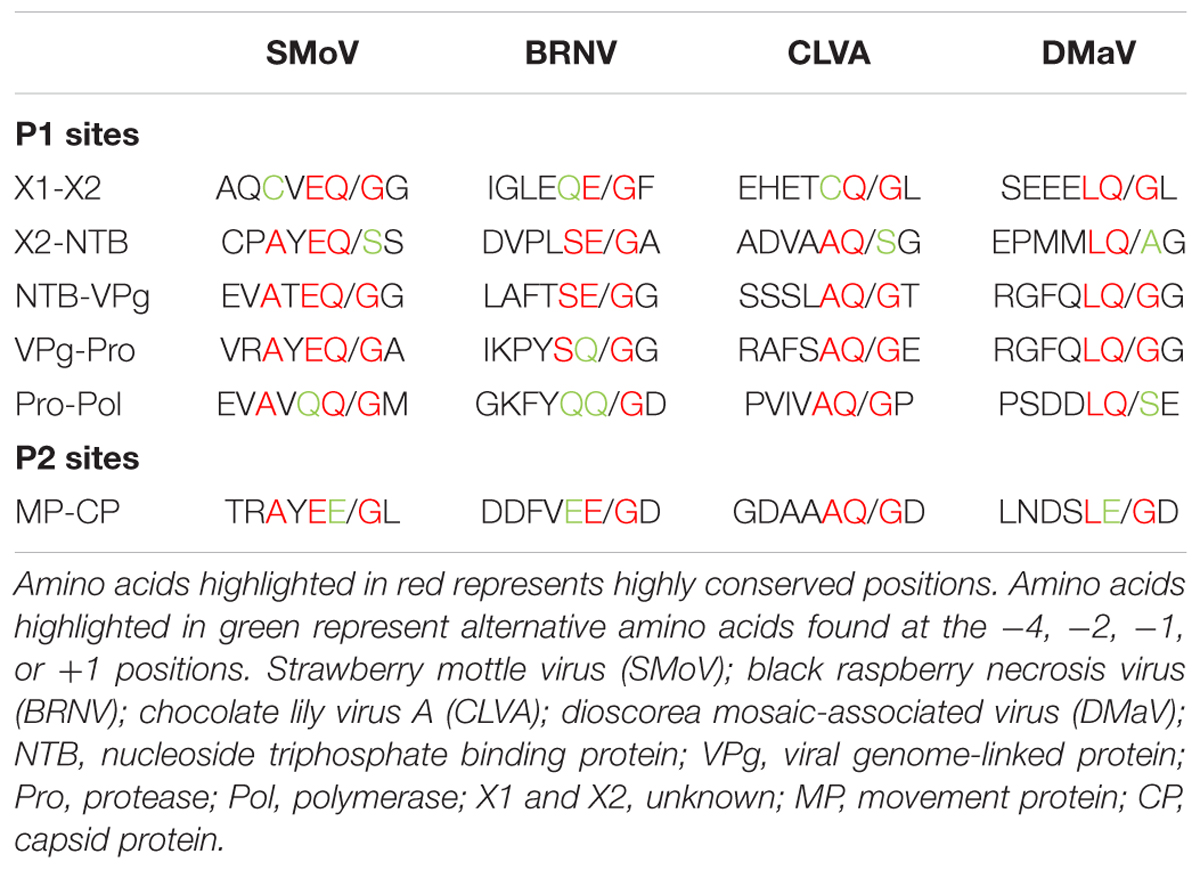

Frontiers | Identification of Cleavage Sites Recognized by the 3C-Like Cysteine Protease within the Two Polyproteins of Strawberry Mottle Virus
Rhinovirus 3C Protease Facilitates Specific Nucleoporin Cleavage and Mislocalisation of Nuclear Proteins in Infected Host Cells | PLOS ONE
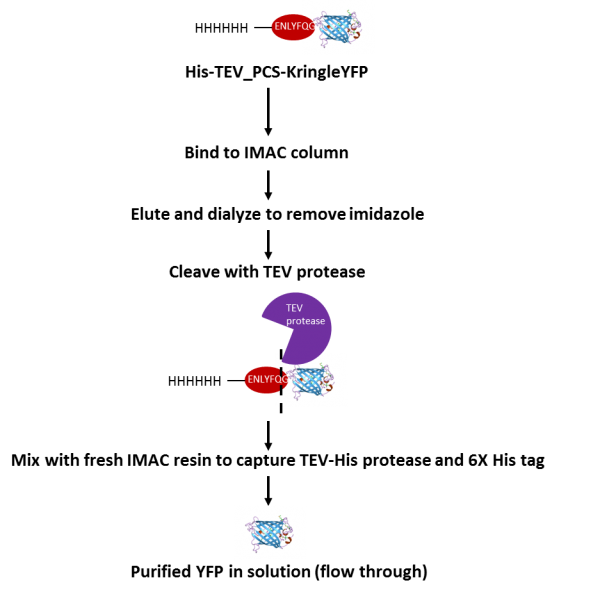
Enzymes - Tobacco Etch Virus (TEV) and Human RhinoVirus (HRV3C) Cysteine Proteases in Vectors | ATUM - ATUM

Directed evolution of the 3C protease from coxsackievirus using a novel fluorescence-assisted intracellular method

Quantitative Analysis of the Substrate Specificity of Human Rhinovirus 3C Protease and Exploration of Its Substrate Recognition Mechanisms | ACS Chemical Biology
Activity of the Human Rhinovirus 3C Protease Studied in Various Buffers, Additives and Detergents Solutions for Recombinant Protein Production | PLOS ONE
![Crystal structure of the 3C protease from Southern African Territories type 2 foot-and-mouth disease virus [PeerJ] Crystal structure of the 3C protease from Southern African Territories type 2 foot-and-mouth disease virus [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2016/1964/1/fig-1-full.png)