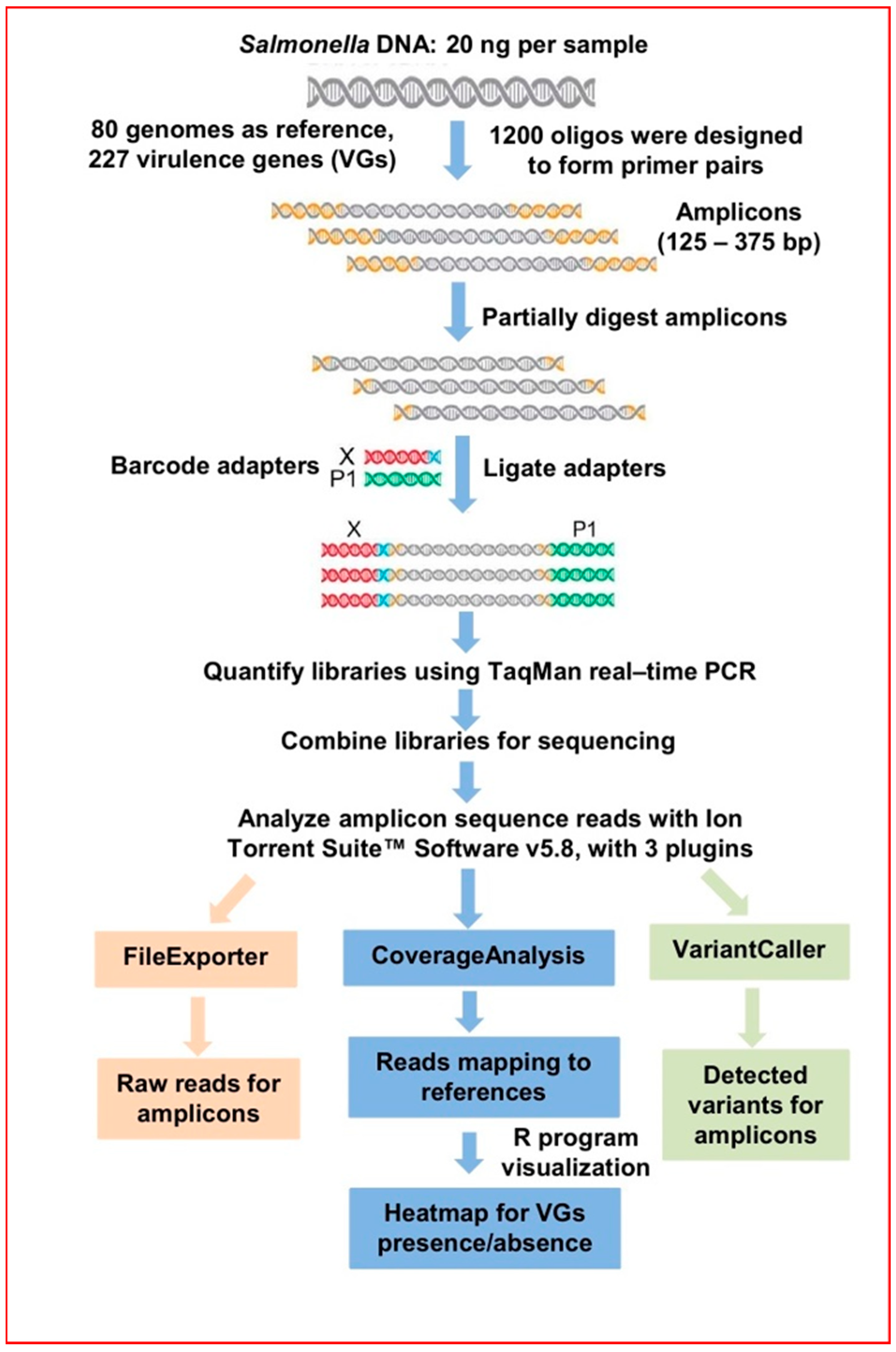
Microorganisms | Free Full-Text | Application of a High-Throughput Targeted Sequence AmpliSeq Procedure to Assess the Presence and Variants of Virulence Genes in Salmonella
Bacterial community structure at the amplicon sequence variant (ASV)... | Download Scientific Diagram
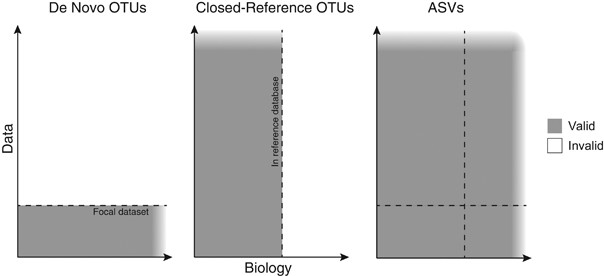
Exact sequence variants should replace operational taxonomic units in marker-gene data analysis | The ISME Journal
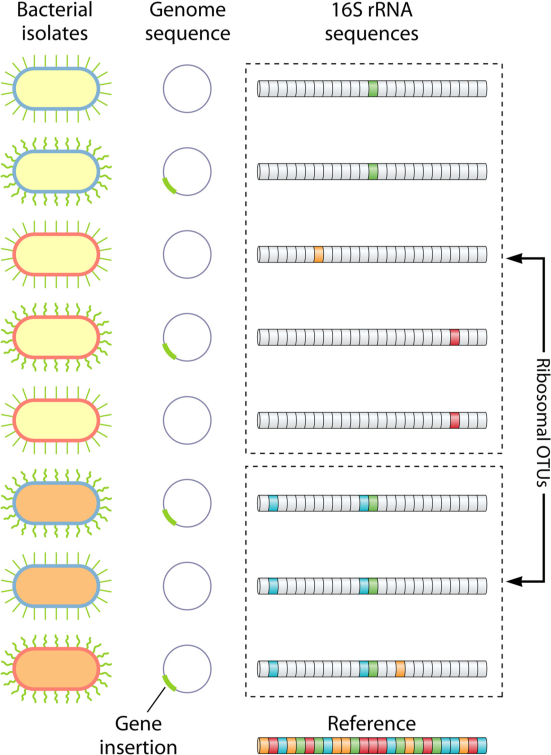
Amplicon sequence variants and bias with Benjamin Callahan — the bioinformatics chat, a podcast about bioinformatics

Population admixtures in medaka inferred by multiple arbitrary amplicon sequencing | Scientific Reports

Frontiers | Cascabel: A Scalable and Versatile Amplicon Sequence Data Analysis Pipeline Delivering Reproducible and Documented Results

Removal of rare amplicon sequence variants from 16S rRNA gene sequence surveys biases the interpretation of community structure data | bioRxiv

evSeq: Cost-Effective Amplicon Sequencing of Every Variant in a Protein Library | ACS Synthetic Biology

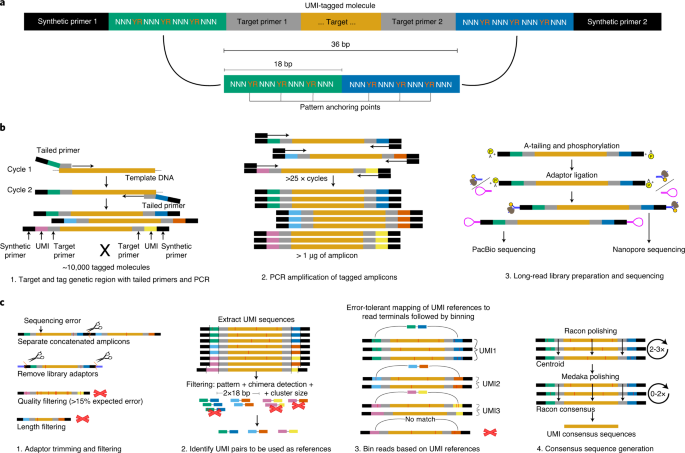
Morten Kam Dahl Dueholm on Twitter: "A new era for 16S rRNA gene amplicon sequencing with near-perfect, full-length amplicon sequence variant (FL-ASV) resolved reference databases. Check out our newly published article in @



![ASV] Amplicon Sequence Variant (ASV)의 특징과 Operational Taxonomic Unit (OTU)와의 차이점 ASV] Amplicon Sequence Variant (ASV)의 특징과 Operational Taxonomic Unit (OTU)와의 차이점](https://blog.kakaocdn.net/dn/Y81DL/btrc6Svly5R/J8lqqEjUQpoPKHawVy0ic0/img.png)