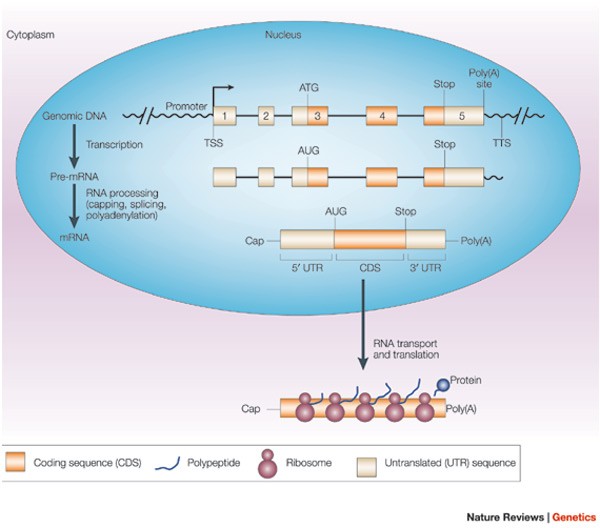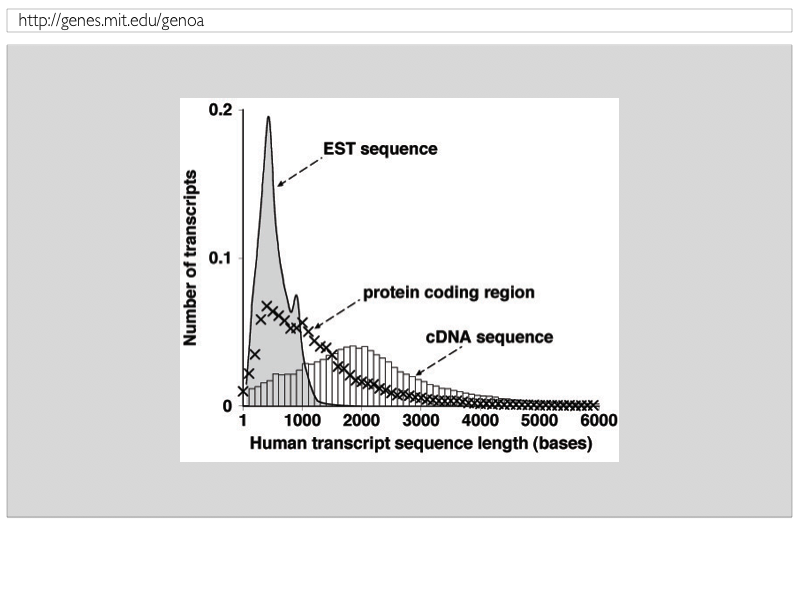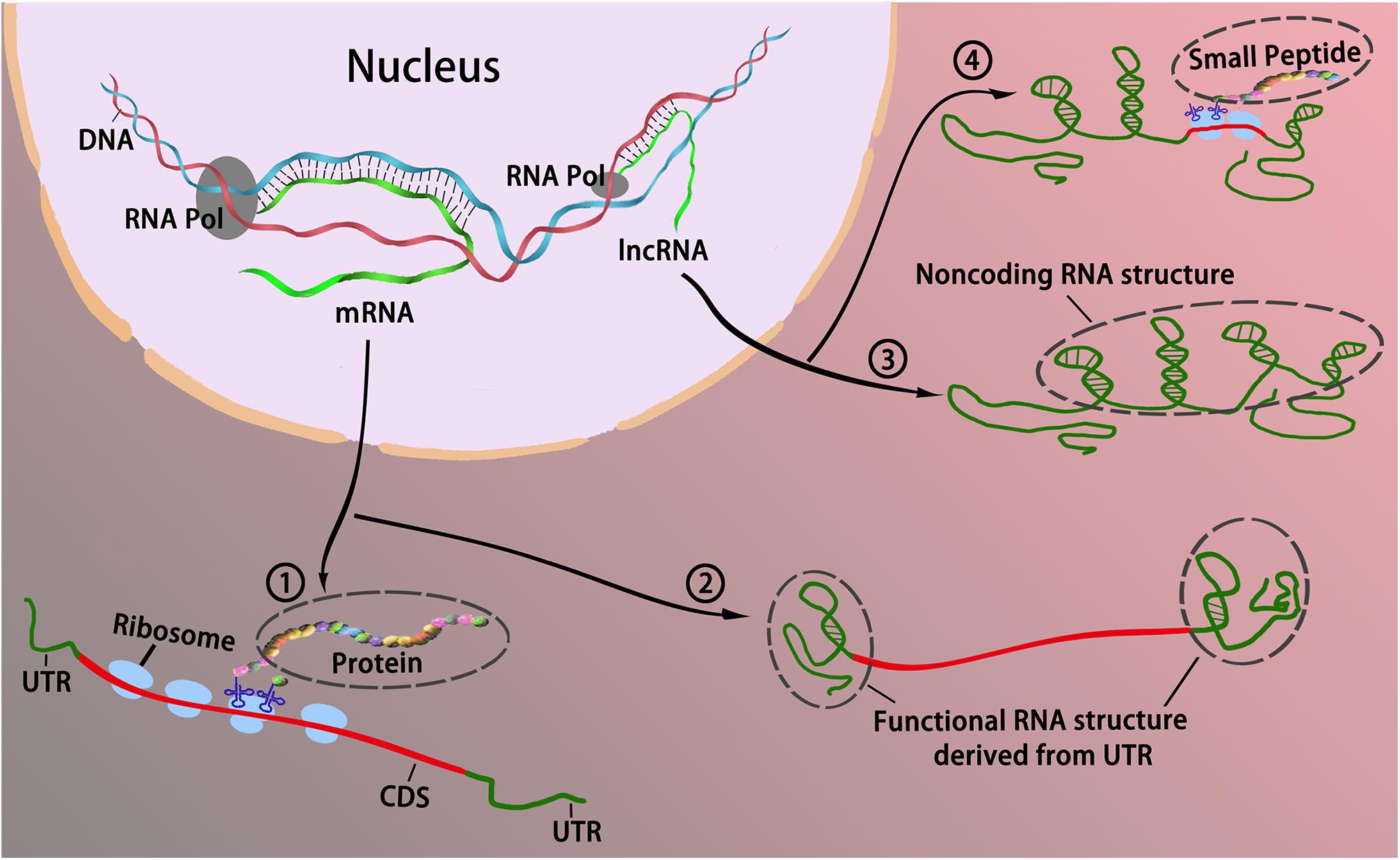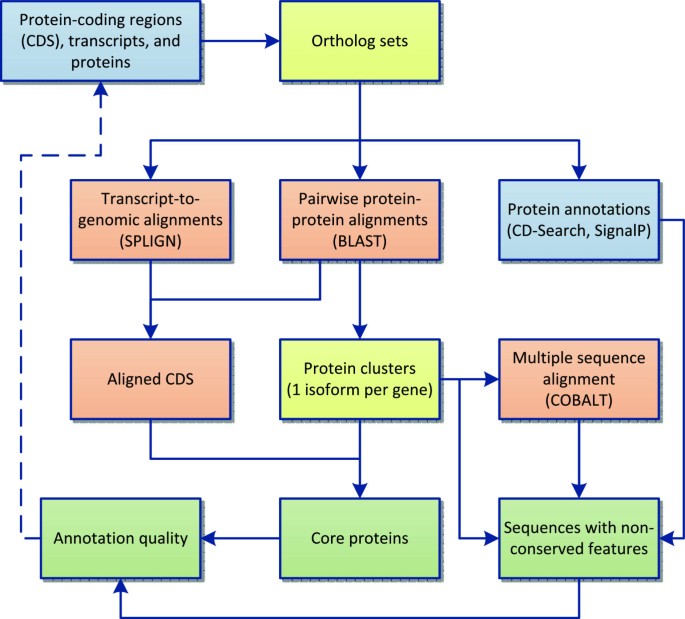
Comparison of RefSeq protein-coding regions in human and vertebrate genomes | BMC Genomics | Full Text

The consensus coding sequence (CCDS) project: Identifying a common protein-coding gene set for the human and mouse genomes
![Open Reading Frame [ORF] - Definition, Role in Protein Synthesis and difference between ORF and CDS - YouTube Open Reading Frame [ORF] - Definition, Role in Protein Synthesis and difference between ORF and CDS - YouTube](https://i.ytimg.com/vi/1T1S6NgECA0/mqdefault.jpg)
Open Reading Frame [ORF] - Definition, Role in Protein Synthesis and difference between ORF and CDS - YouTube

Geno2proteo, a Tool for Batch Retrieval of DNA and Protein Sequences from Any Genomic or Protein Regions

Exploring a Nucleotide Sequence Using the Sequence Viewer App - MATLAB & Simulink - MathWorks Deutschland
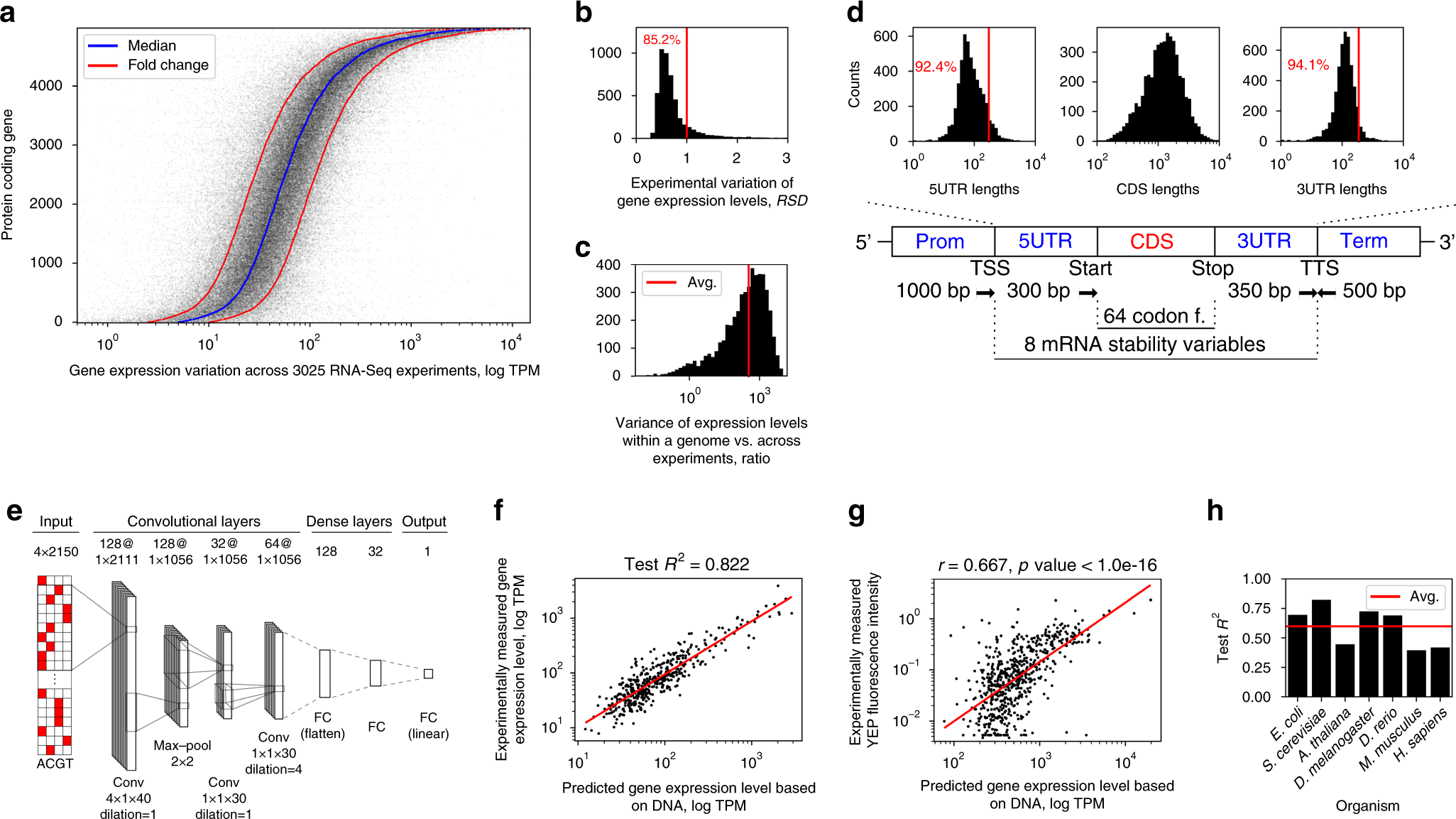
Deep learning suggests that gene expression is encoded in all parts of a co-evolving interacting gene regulatory structure | Nature Communications