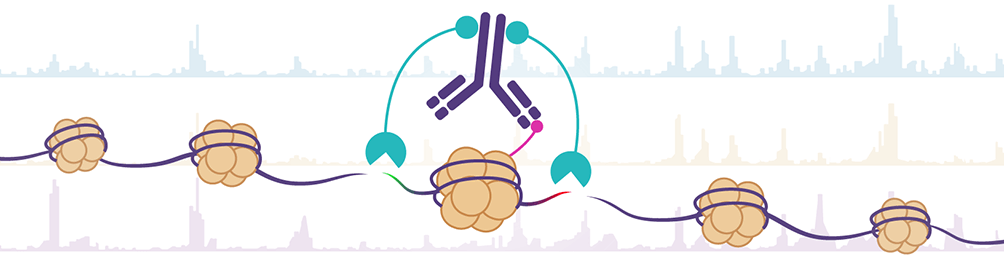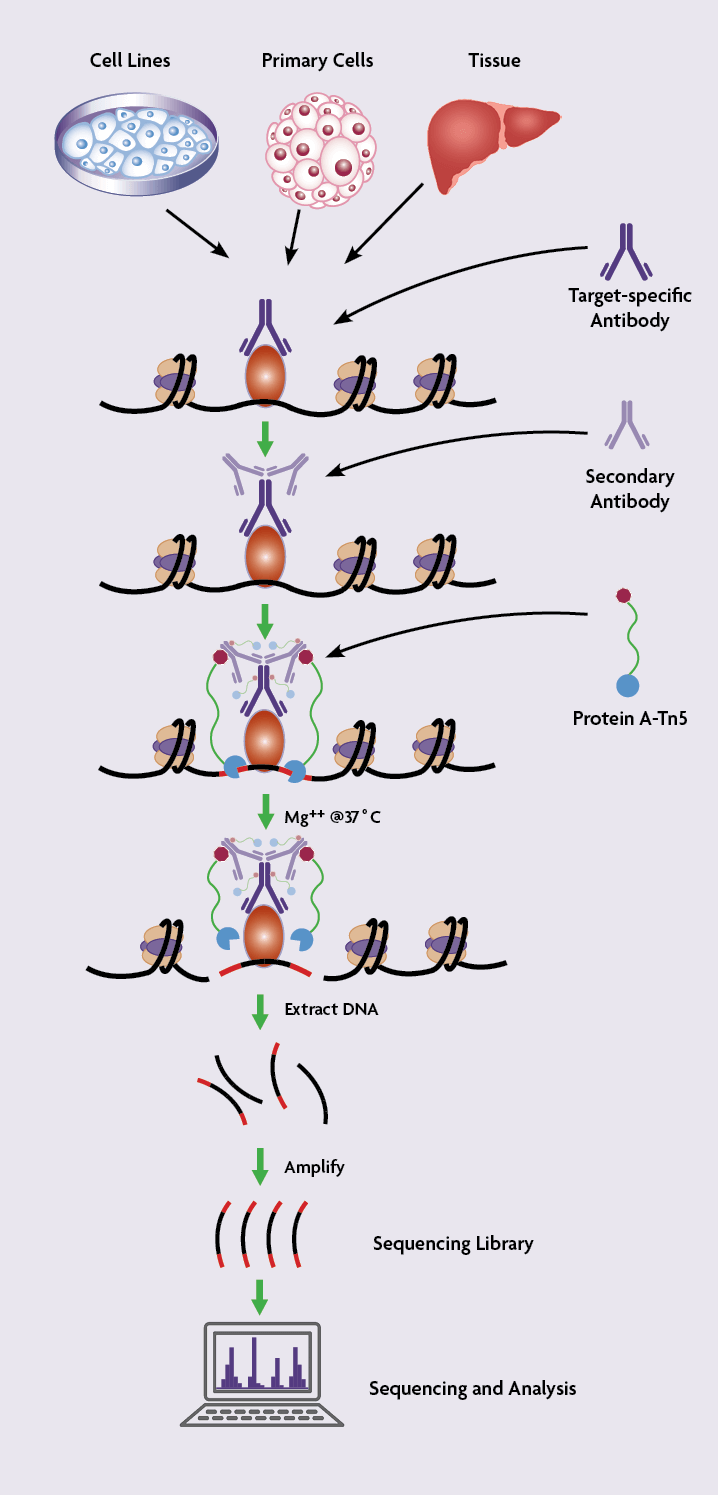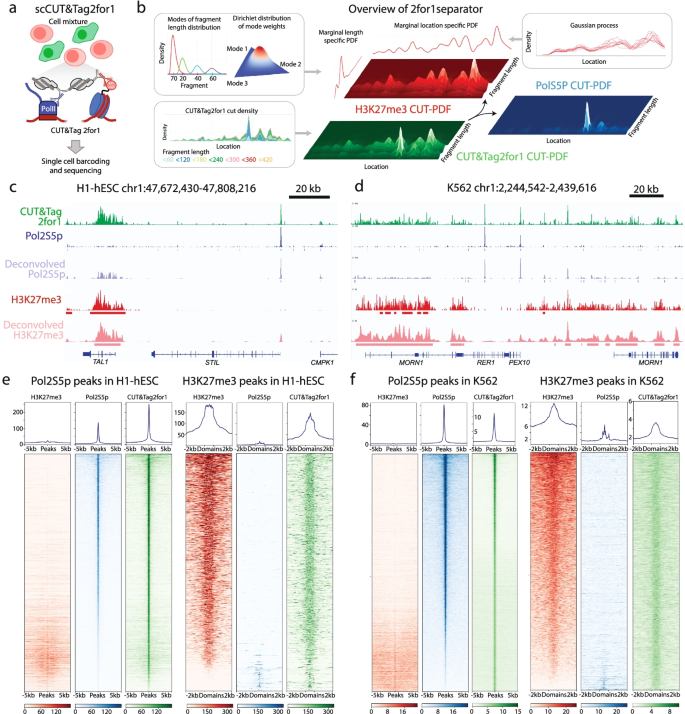
CUT&Tag2for1: a modified method for simultaneous profiling of the accessible and silenced regulome in single cells | Genome Biology | Full Text

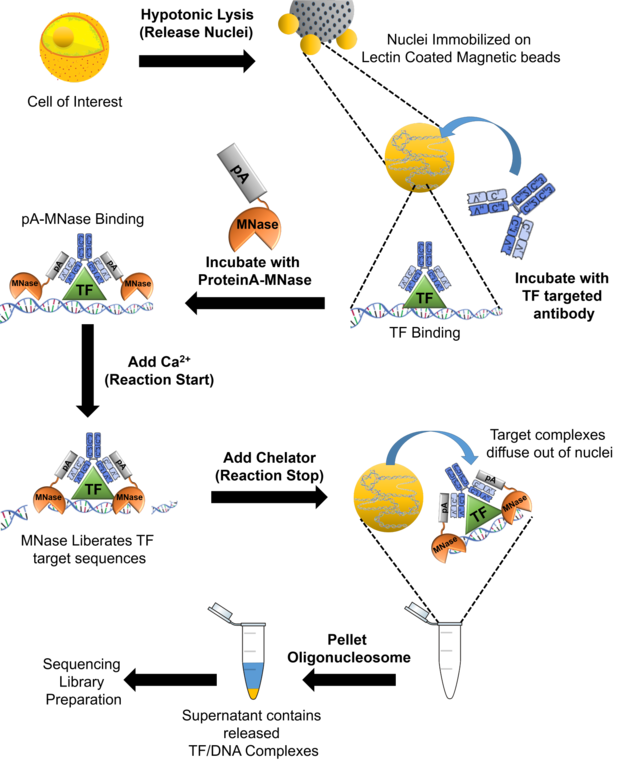
Overview of TIP-seq, a robust low-cell mapping method that combines... | Download Scientific Diagram
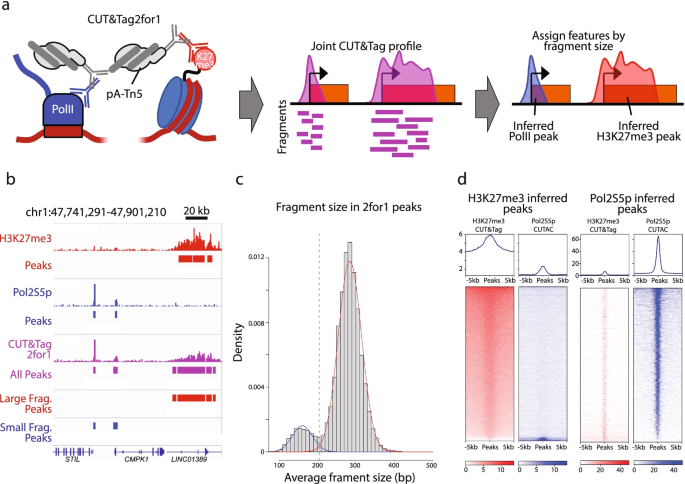
CUT&Tag2for1: a modified method for simultaneous profiling of the accessible and silenced regulome in single cells | Genome Biology | Full Text

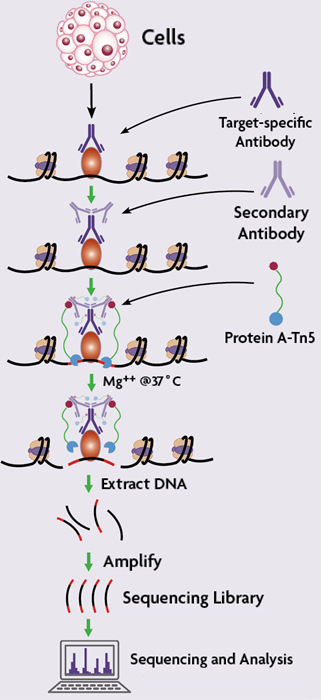
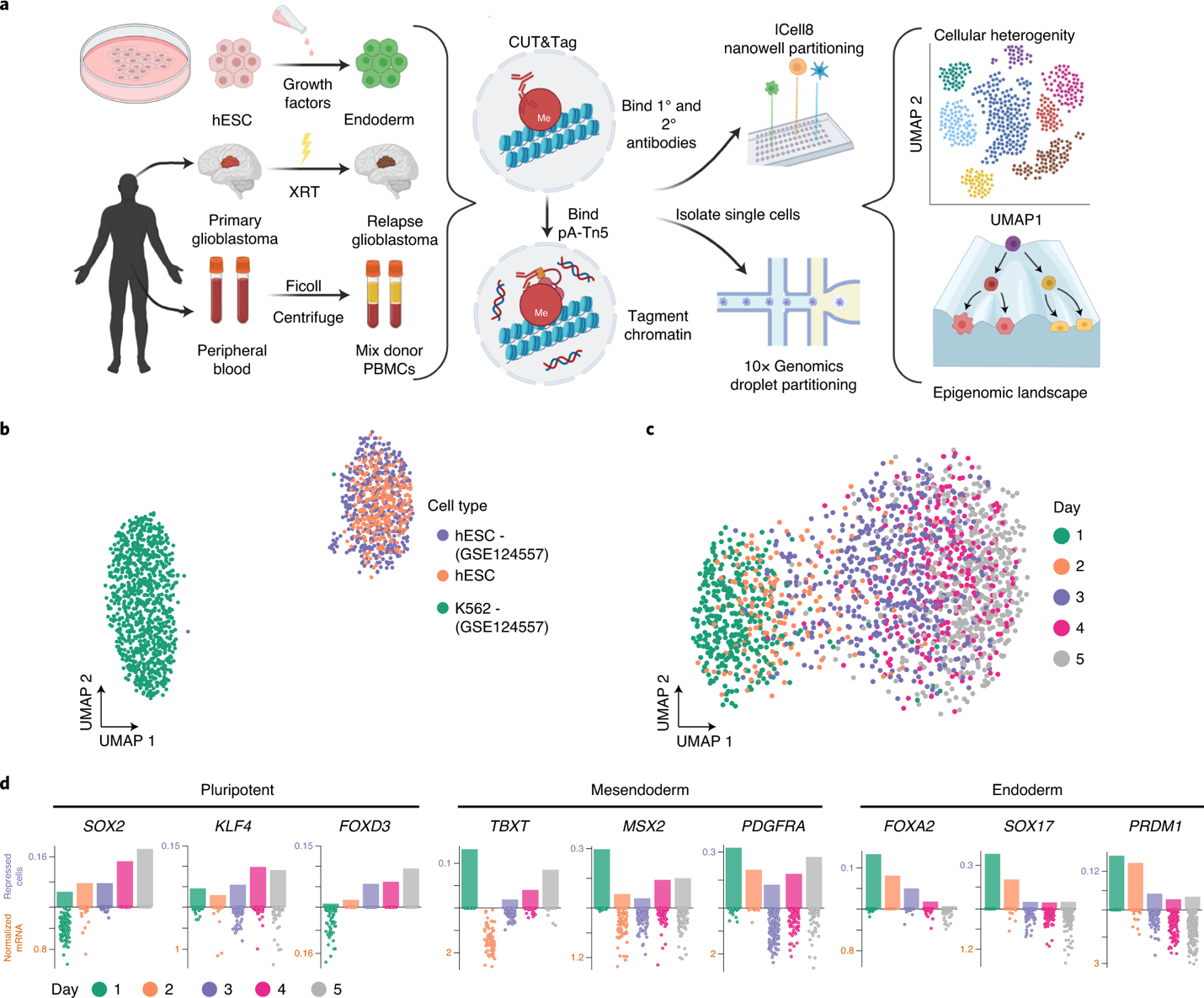
CUT&Tag for efficient epigenomic profiling of small samples and single cells | Nature Communications

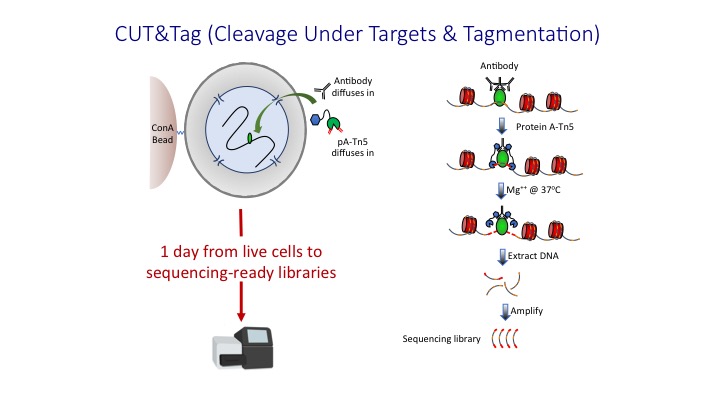
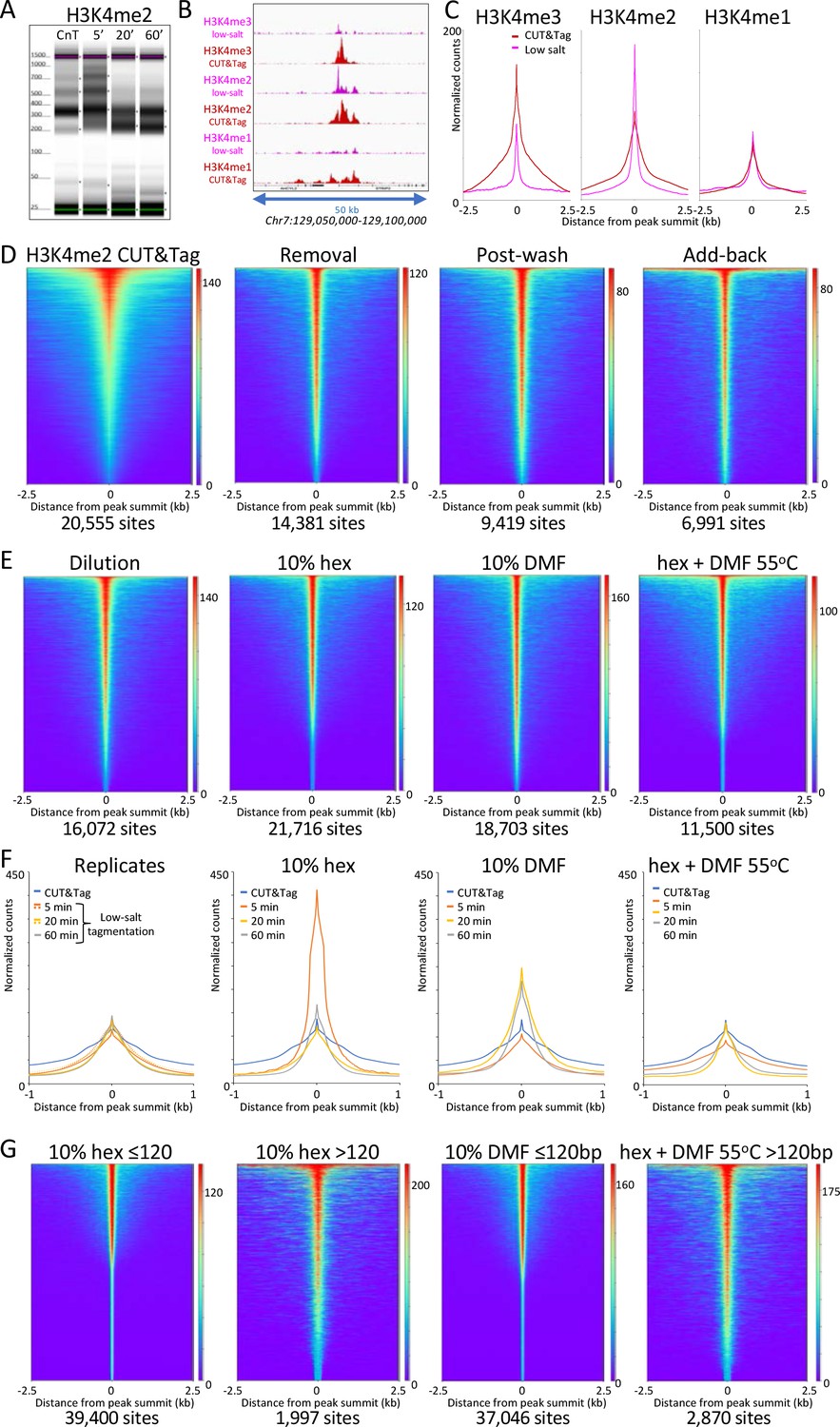
CUT&Tag with low-salt tagmentation (CUTAC). Steps in gray are lab-based... | Download Scientific Diagram
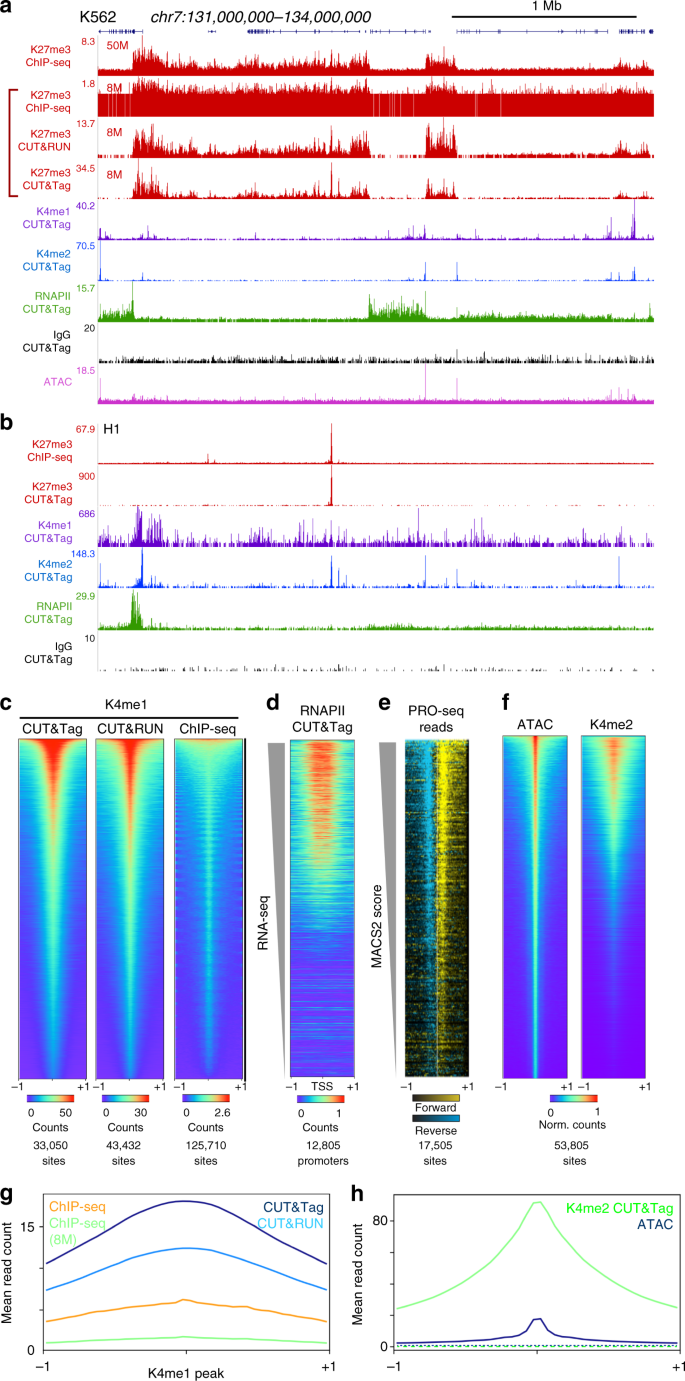
CUT&Tag for efficient epigenomic profiling of small samples and single cells | Nature Communications