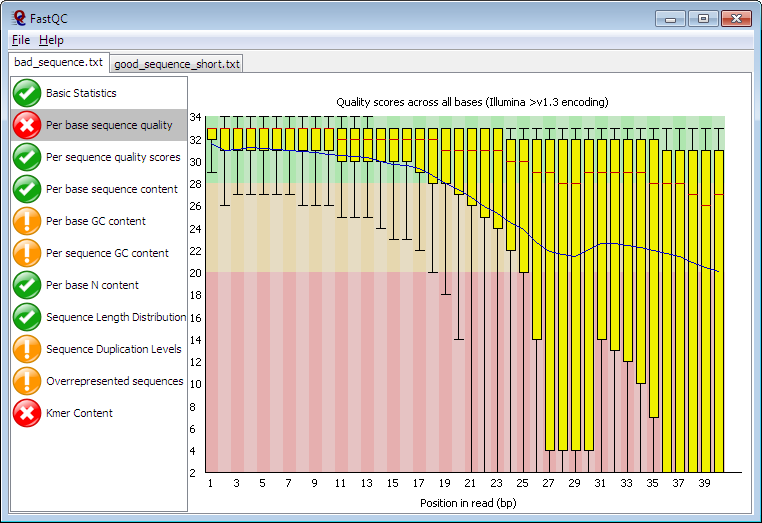
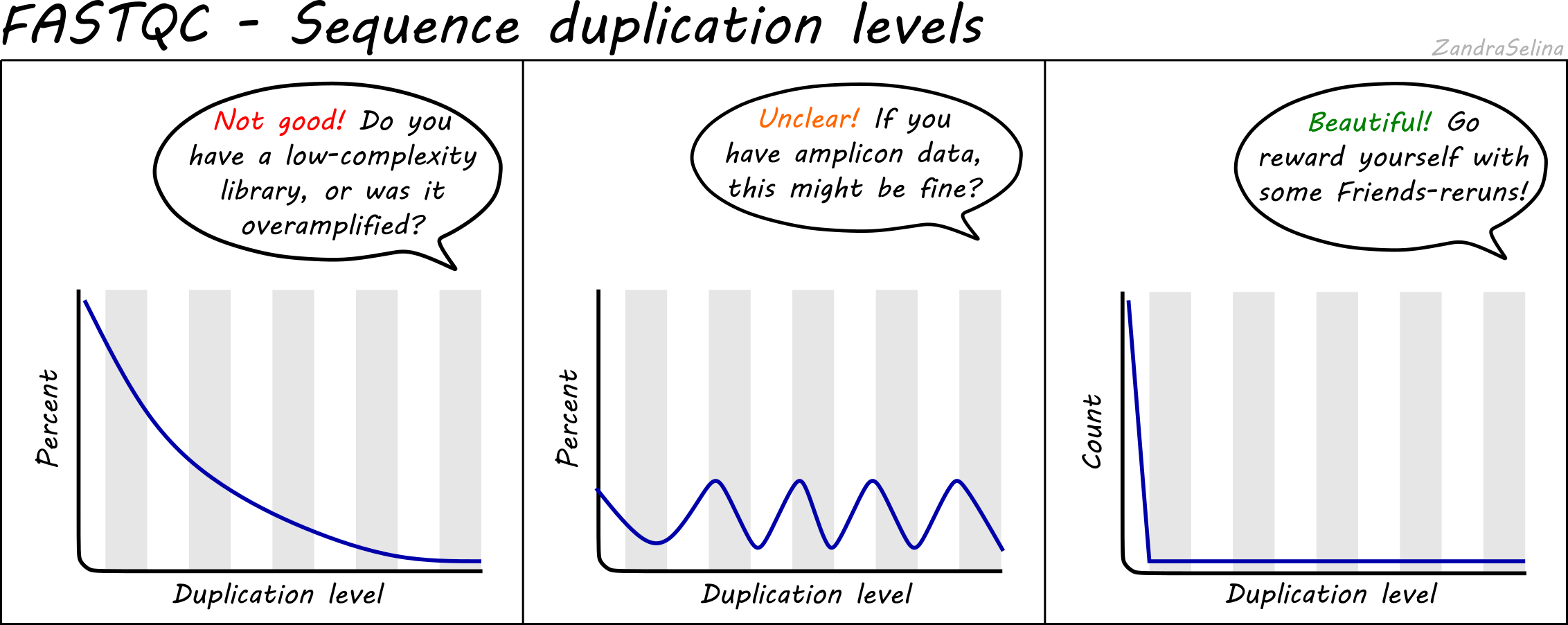
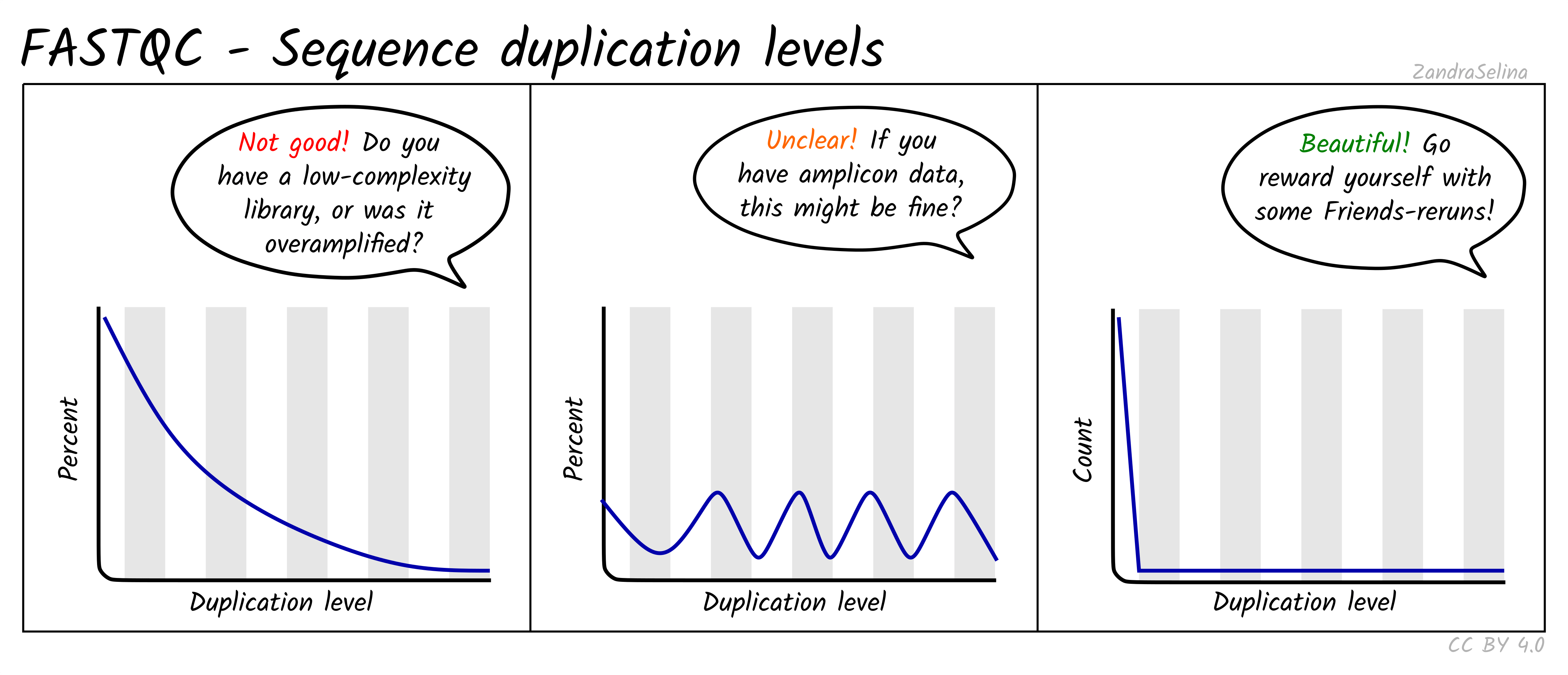
FASTQ Quality Assurance Tools - Bioinformatics Team (BioITeam) at the University of Texas - UT Austin Wikis
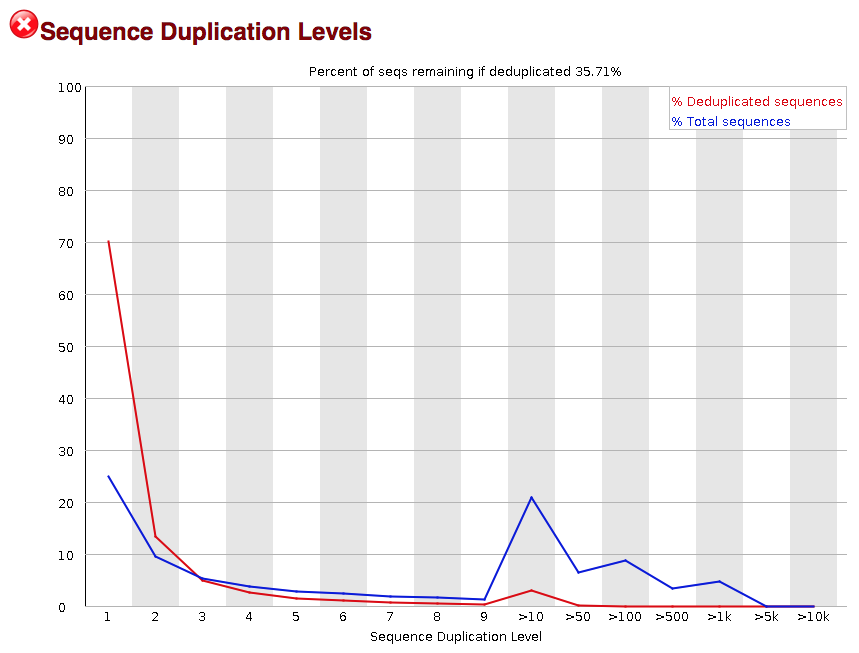
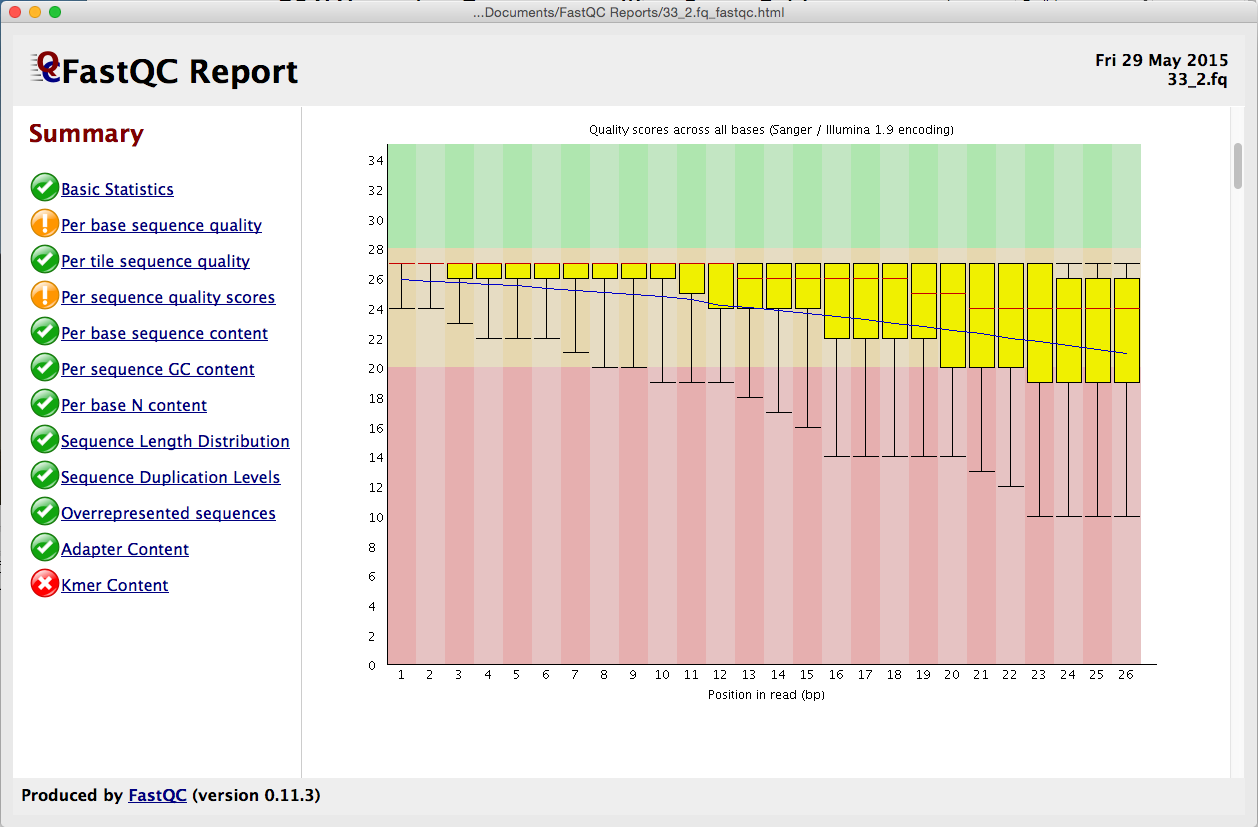
Quality control: Assessing FASTQC results | Introduction to RNA-Seq using high-performance computing - ARCHIVED

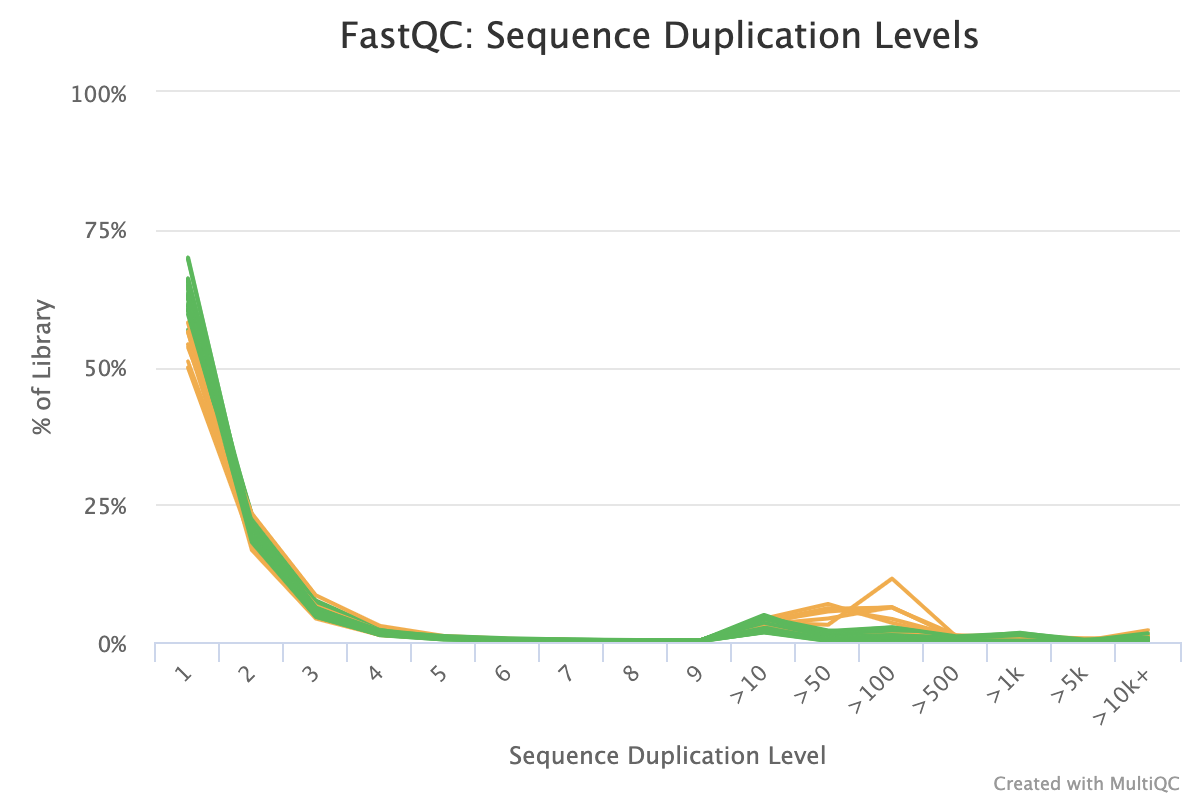
FastQC: Sequence Duplication Levels interactive dialog indexing error · Issue #1092 · ewels/MultiQC · GitHub
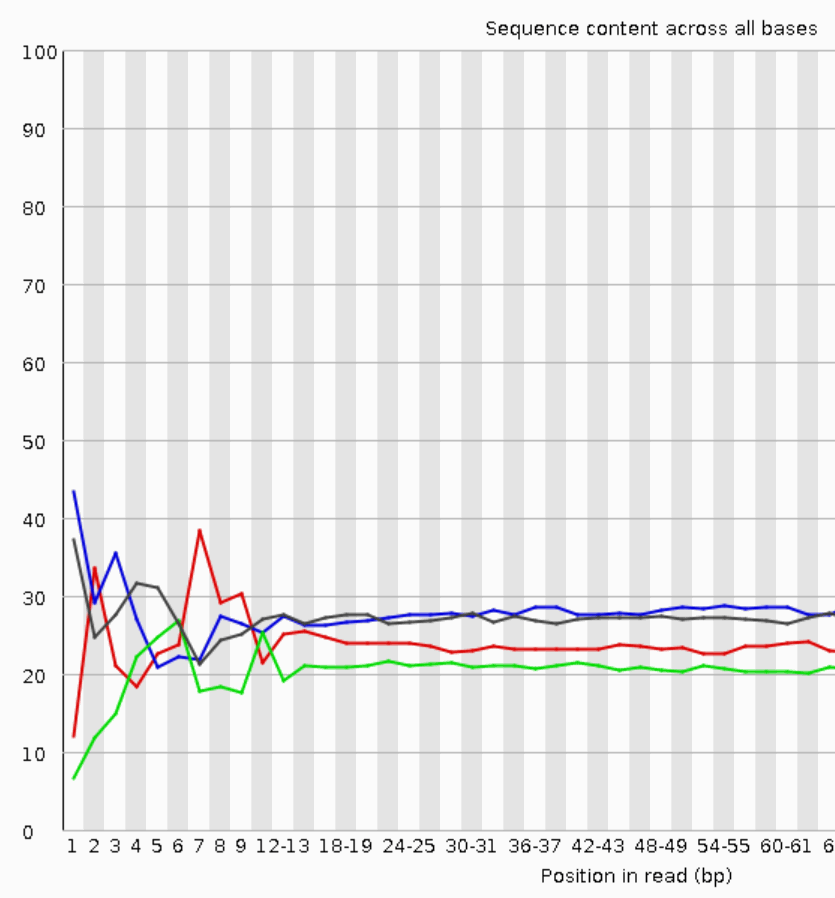
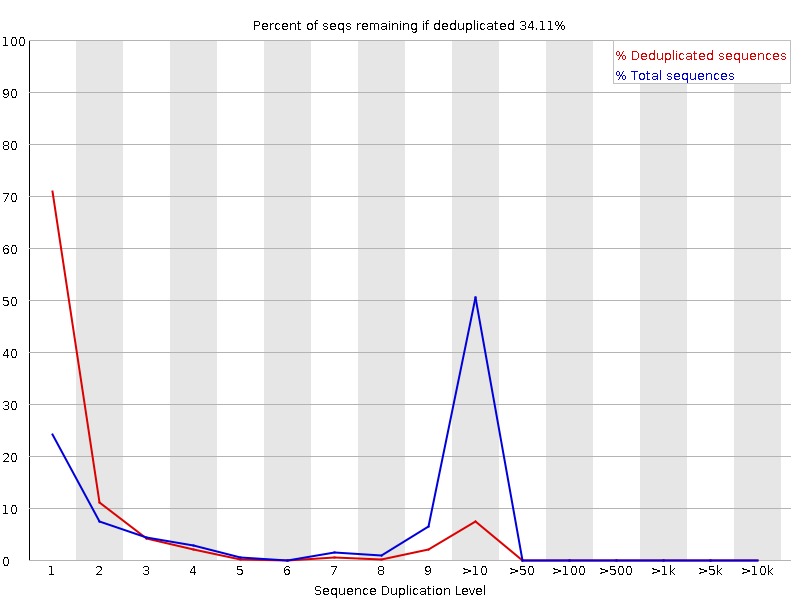
Why does FASTQC show unexpectedly high sequence duplication levels (PCR-duplicates)? | DNA Technologies Core

Duplication rate differs up to 30% from that of Fastqc for single end reads · Issue #290 · OpenGene/fastp · GitHub
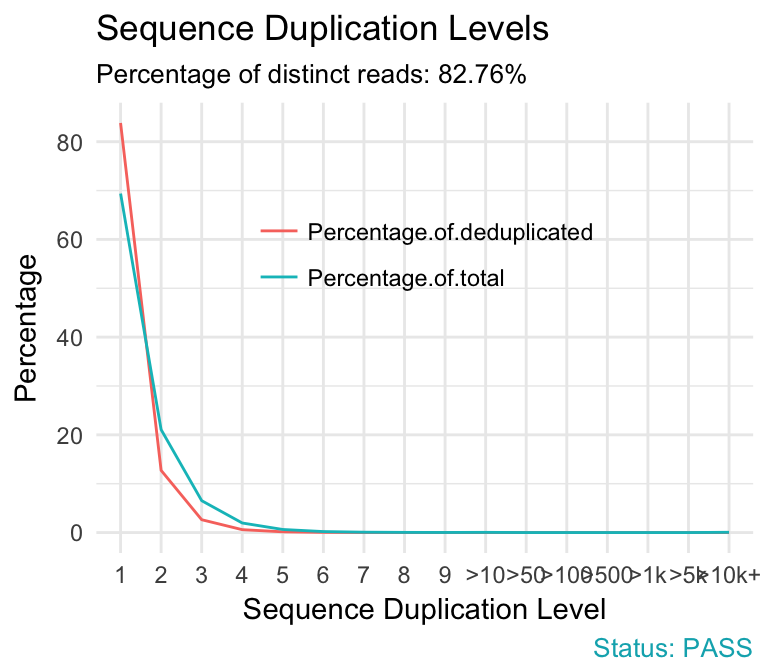
fastqcr: An R Package Facilitating Quality Controls of Sequencing Data for Large Numbers of Samples | R-bloggers