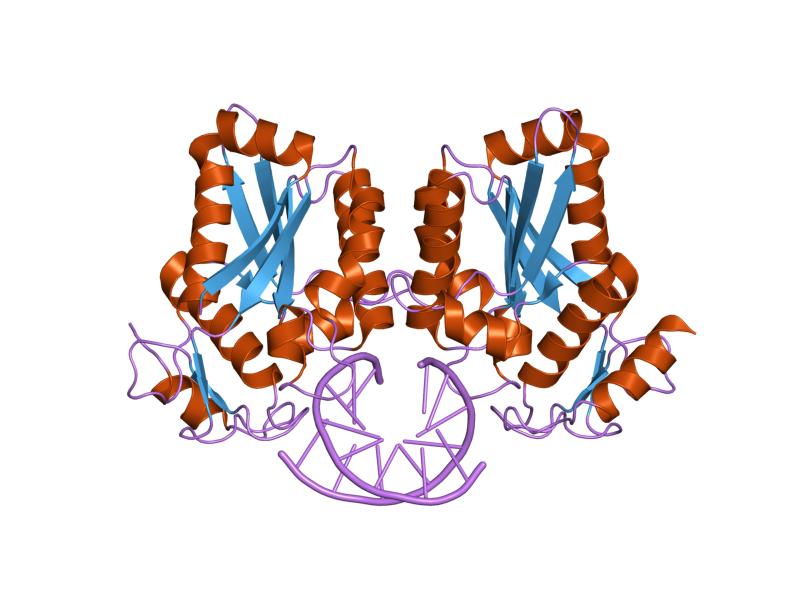
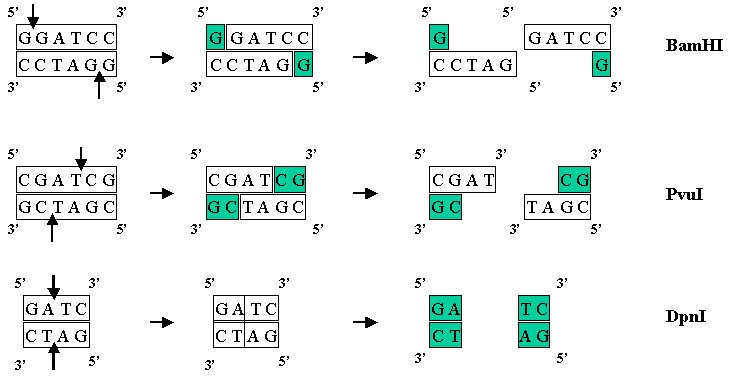Implications for Switching Restriction Enzyme Specificities from the Structure of BstYI Bound to a BglII DNA Sequence - ScienceDirect
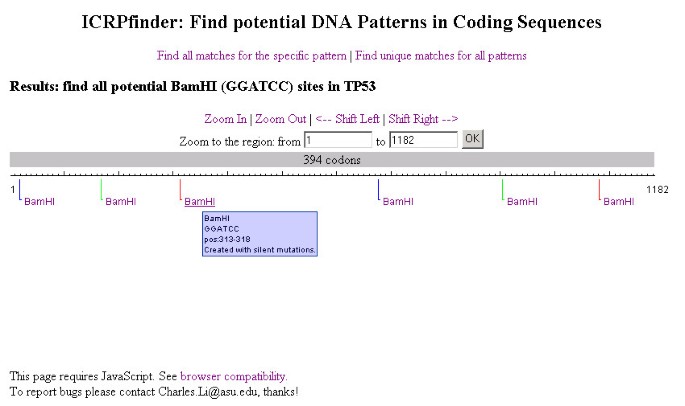
ICRPfinder: a fast pattern design algorithm for coding sequences and its application in finding potential restriction enzyme recognition sites | BMC Bioinformatics | Full Text
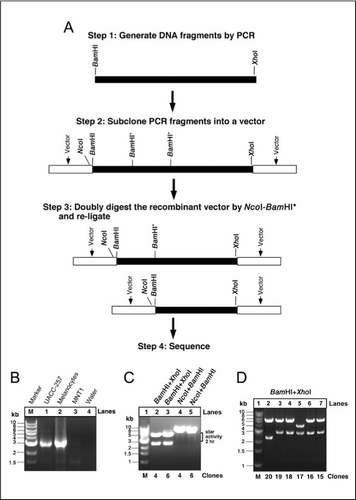
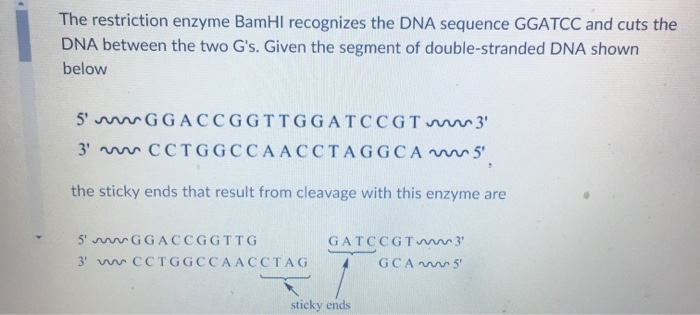
Useful tool to generate unidirectional deletion vectors by utilizing the star activity of BamHI in an NcoI-BamHI-XhoI cassette | BioTechniques
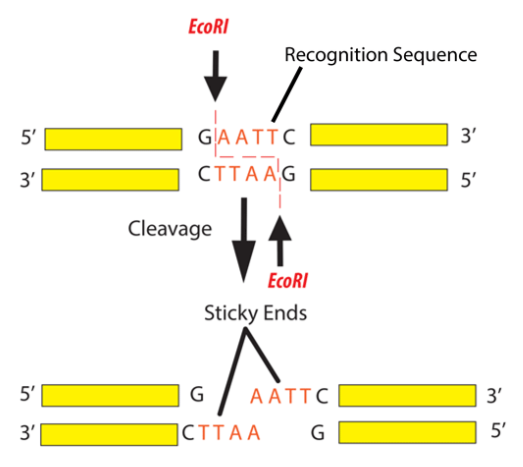
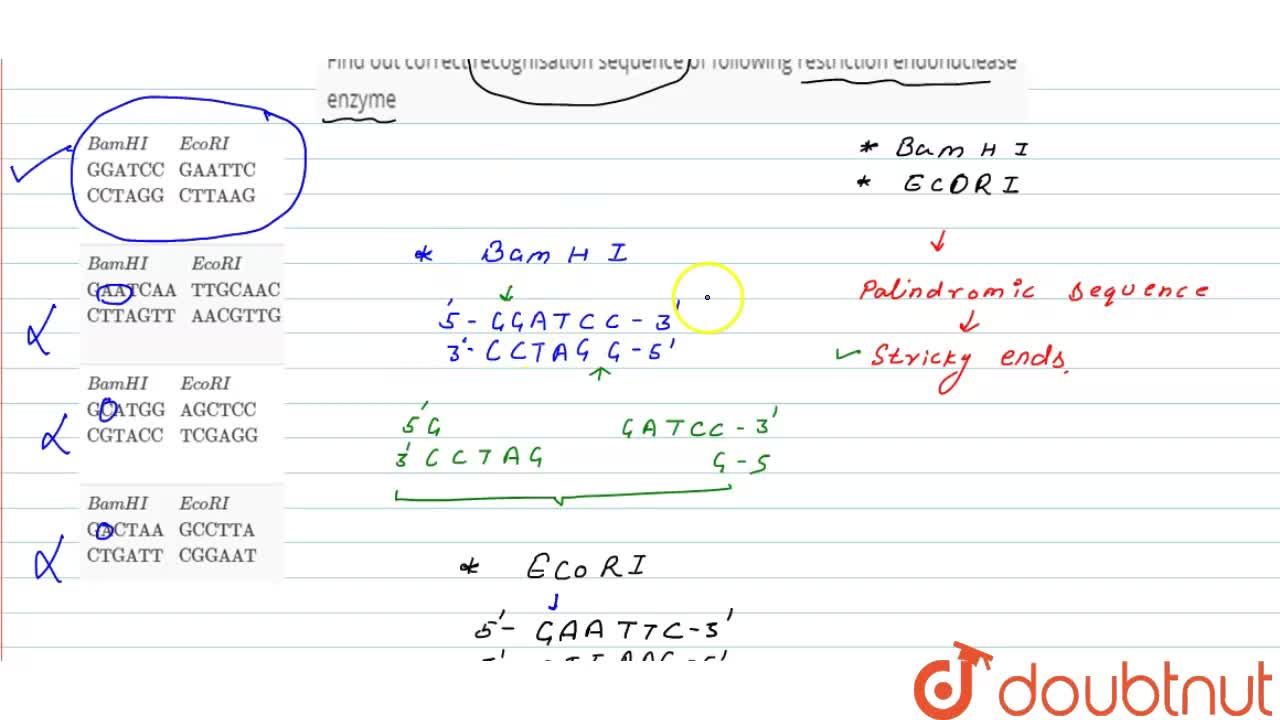
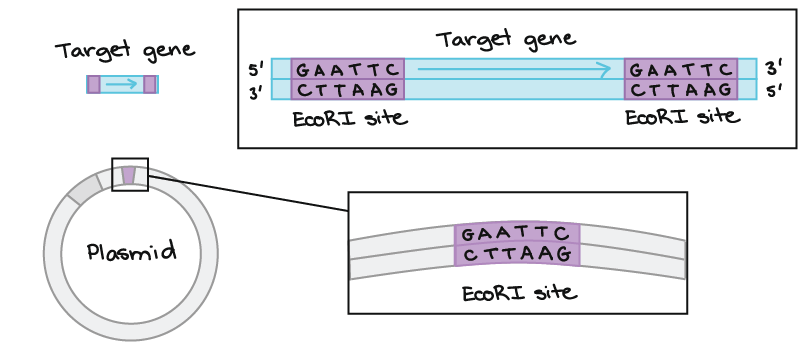
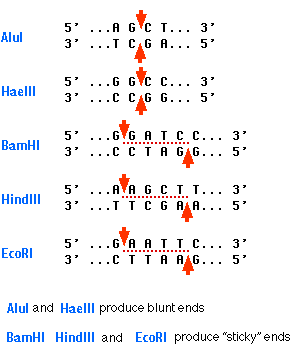
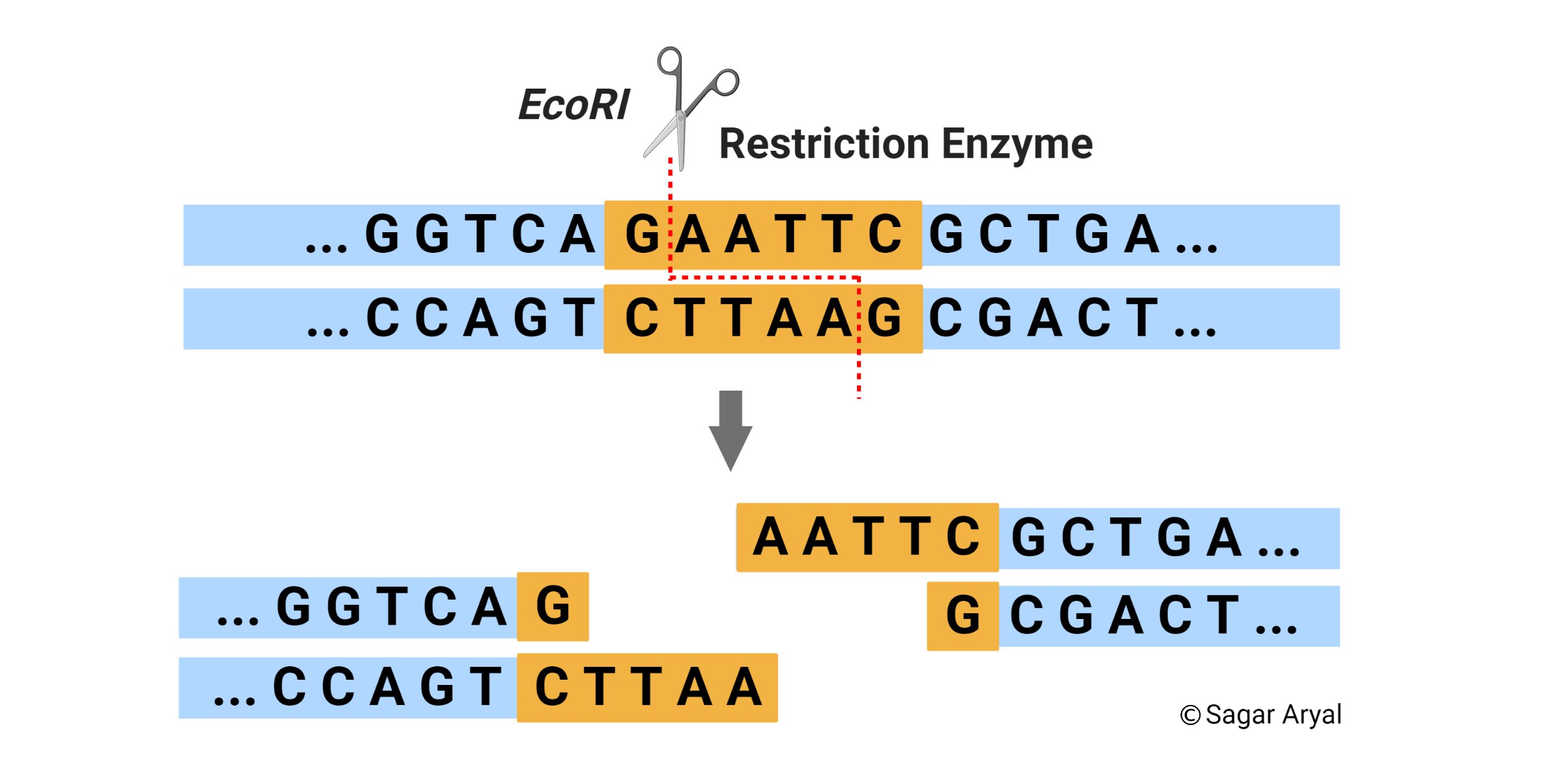
GAATTC is the recognition site for which of the following restrictions endonuclease? a. Hind IIIb. EcoR Ic. Bam IId. Hae III
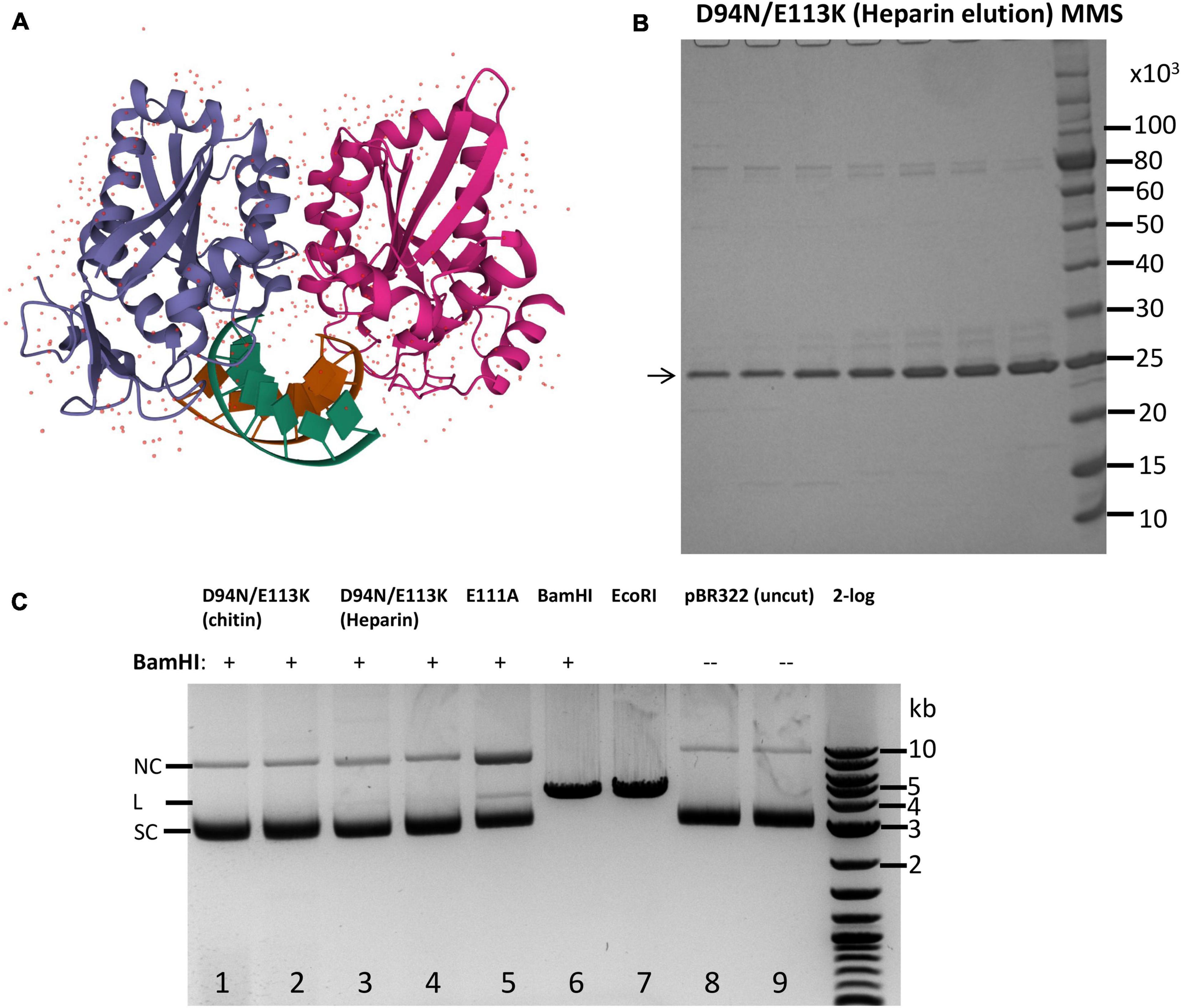
Frontiers | Engineering Infrequent DNA Nicking Endonuclease by Fusion of a BamHI Cleavage-Deficient Mutant and a DNA Nicking Domain