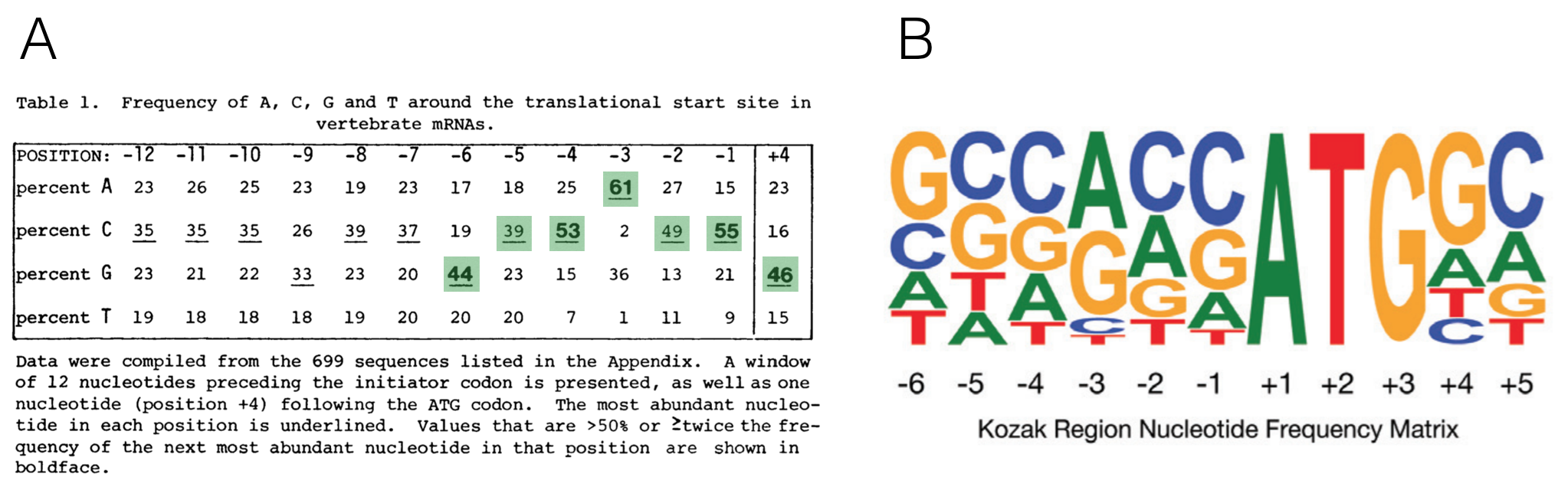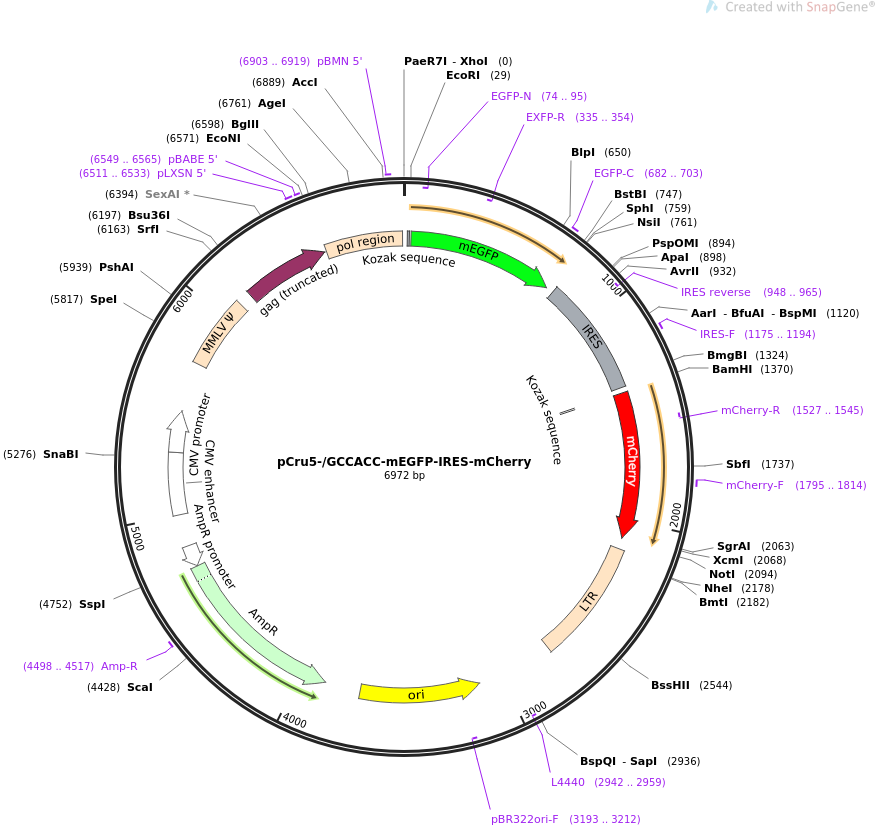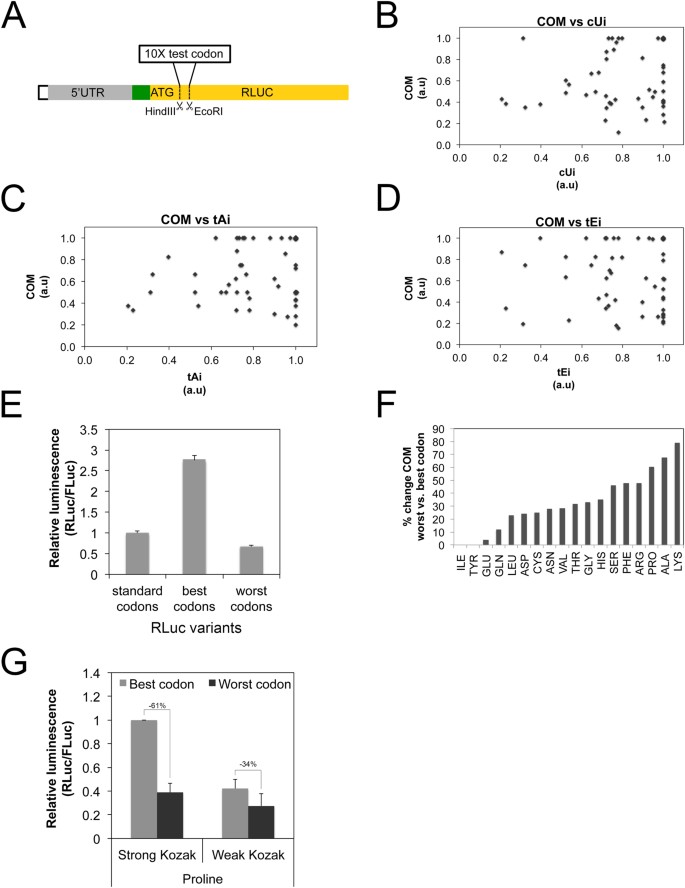
Changes in global translation elongation or initiation rates shape the proteome via the Kozak sequence | Scientific Reports

Quantitative analysis of mammalian translation initiation sites by FACS‐seq | Molecular Systems Biology
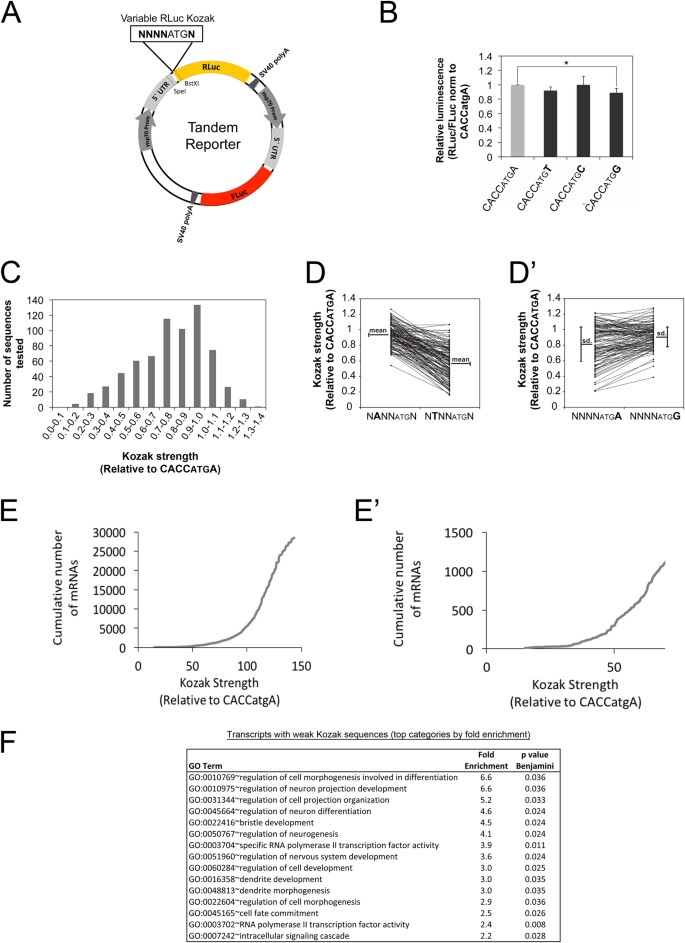
Changes in global translation elongation or initiation rates shape the proteome via the Kozak sequence | Scientific Reports
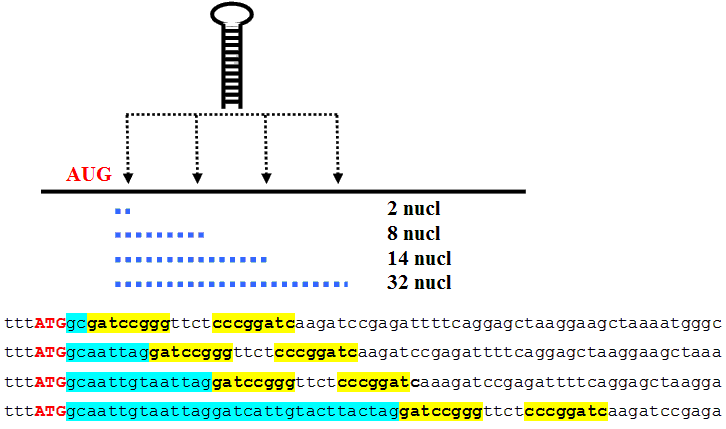
AUG_hairpin: program for prediction of a downstream hairpin potentially increasing initiation of translation at start AUG codon in a suboptimal context.

Bioinformatic analyses of mammalian 5'-UTR sequence properties of mRNAs predicts alternative translation initiation sites | BMC Bioinformatics | Full Text

Kozak consensus sequence surrounding start methionin in the 440 cCDS... | Download Scientific Diagram

Enhancement of protein expression in insect cells by a lobster tropomyosin cDNA leader sequence - ScienceDirect
Alternative Transcription Initiation and the AUG Context Configuration Control Dual-Organellar Targeting and Functional Competen

Structural insights into the Kozak sequence interactions within the mammalian late-stage initiation complexes | bioRxiv

Machine learning predicts translation initiation sites in neurologic diseases with expanded repeats | bioRxiv