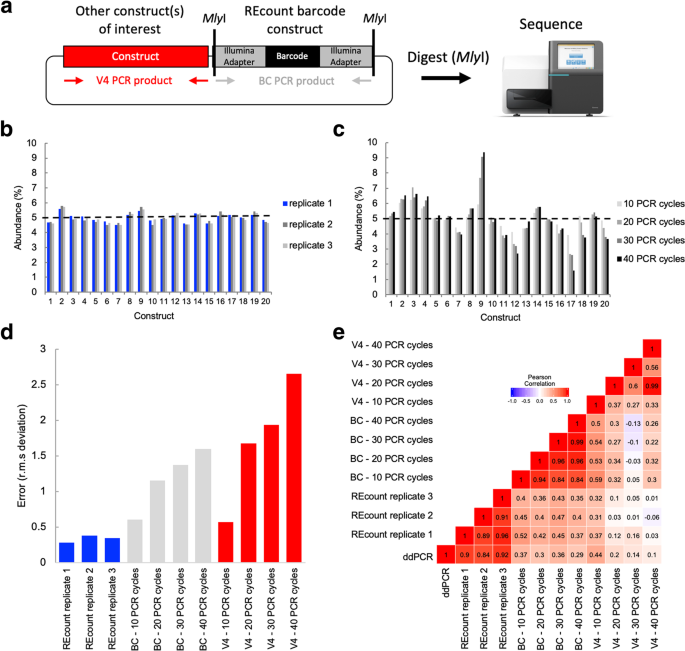
Measuring sequencer size bias using REcount: a novel method for highly accurate Illumina sequencing-based quantification | Genome Biology | Full Text
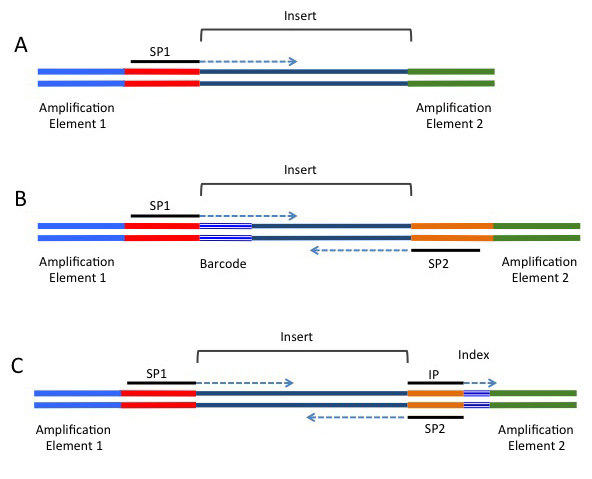
Biotech7005 | The practical material from the course Biotech 7005: Bioinformatics and Systems Modelling

Illumina Genome Analyzer sequencing. Adapter-modified, single-stranded... | Download Scientific Diagram

Adapter trimming: Why are adapter sequences trimmed from only the 3' ends of reads - Illumina Knowledge

Ultrahigh-Throughput Multiplexing and Sequencing of >500-Base-Pair Amplicon Regions on the Illumina HiSeq 2500 Platform | mSystems
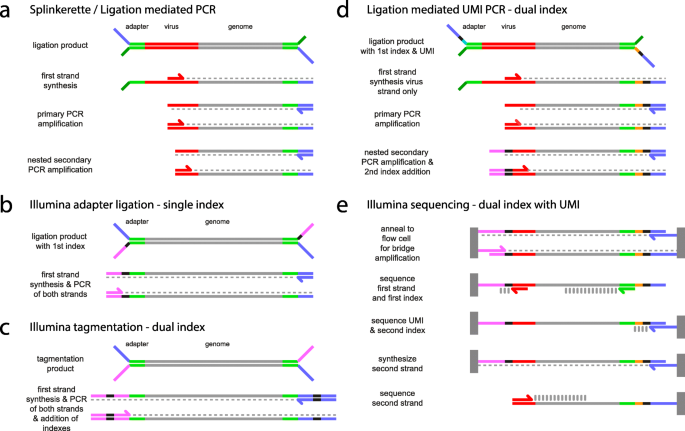
LUMI-PCR: an Illumina platform ligation-mediated PCR protocol for integration site cloning, provides molecular quantitation of integration sites | Mobile DNA | Full Text
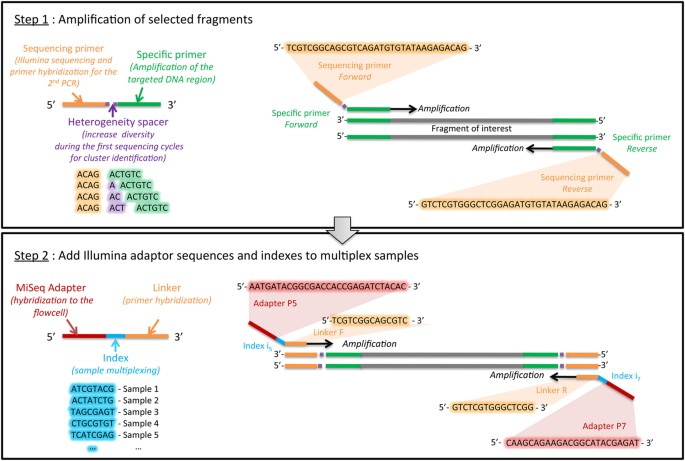
High-throughput sequencing of multiple amplicons for barcoding and integrative taxonomy | Scientific Reports

Universal two-step PCR approach. First-round PCR is performed by using... | Download Scientific Diagram
![Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ] Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7755/1/fig-4-2x.jpg)
Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ]





![Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ] Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7786/1/fig-3-full.png)