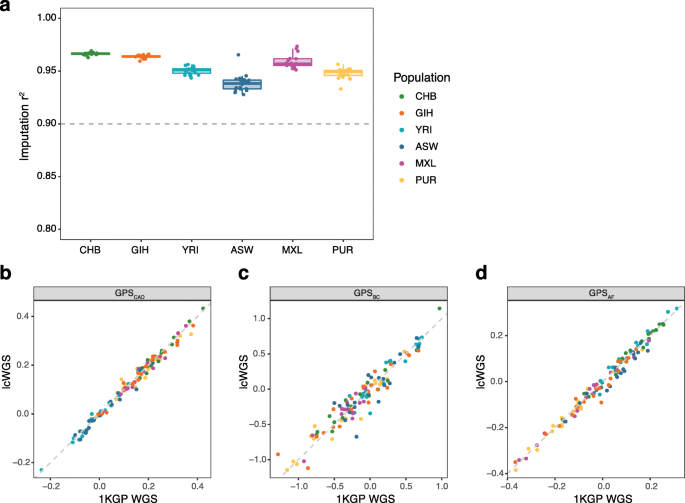
Low coverage whole genome sequencing enables accurate assessment of common variants and calculation of genome-wide polygenic scores | Genome Medicine | Full Text

Visualization of the Chr7q35 deletion using the UCSC genome browser.... | Download Scientific Diagram

Low-coverage whole-genome sequencing of extracellular vesicle-associated DNA in patients with metastatic cancer | Scientific Reports
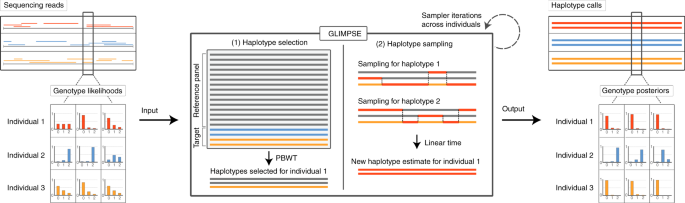
Efficient phasing and imputation of low-coverage sequencing data using large reference panels | Nature Genetics
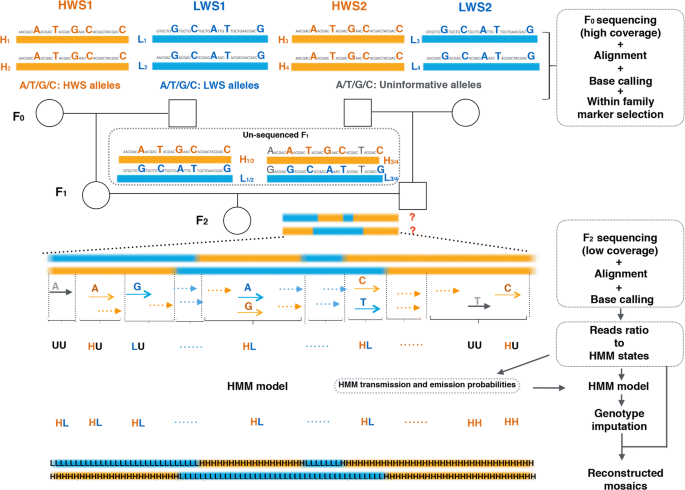
Genotyping by low-coverage whole-genome sequencing in intercross pedigrees from outbred founders: a cost-efficient approach | Genetics Selection Evolution | Full Text
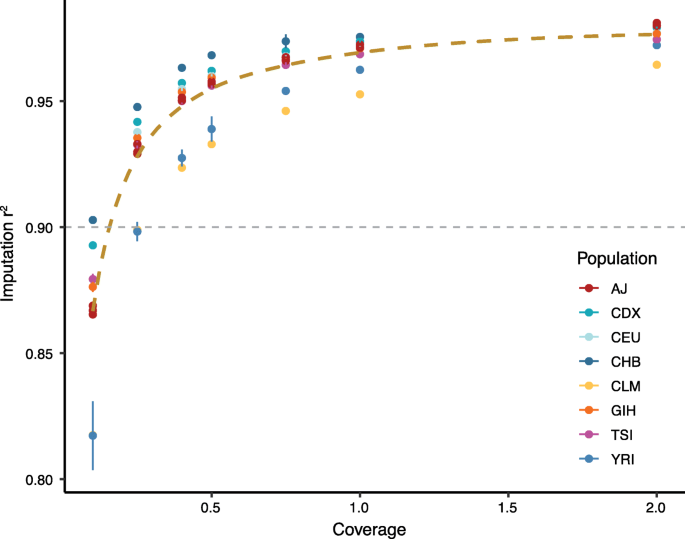
Low coverage whole genome sequencing enables accurate assessment of common variants and calculation of genome-wide polygenic scores | Genome Medicine | Full Text
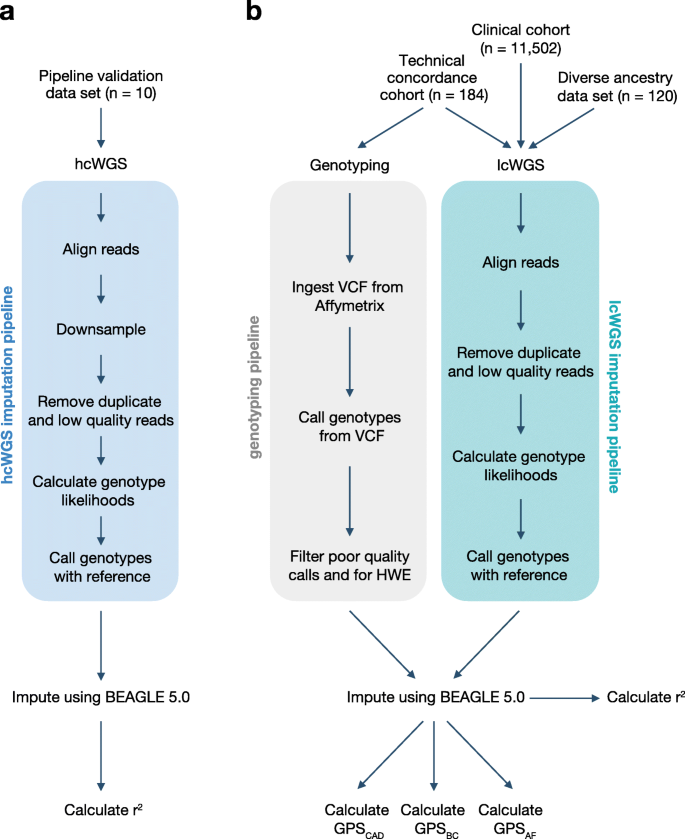
Low coverage whole genome sequencing enables accurate assessment of common variants and calculation of genome-wide polygenic scores | Genome Medicine | Full Text

Low-coverage sequencing cost-effectively detects known and novel variation in underrepresented populations - ScienceDirect

A Highly Scalable Method for Joint Whole-Genome Sequencing and Gene-Expression Profiling of Single Cells - ScienceDirect