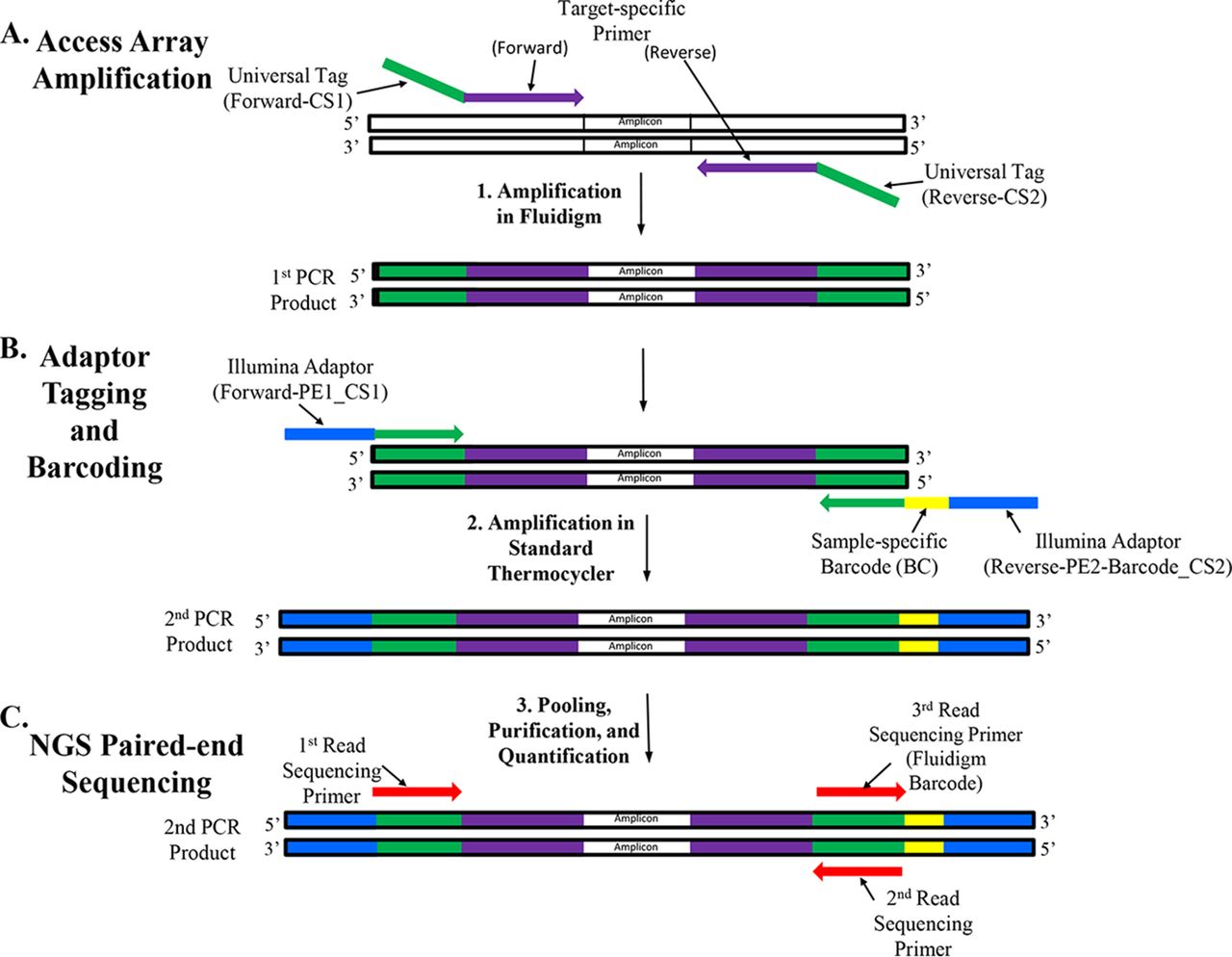
Illumina MiSeq™ system | The Genomics Research Unit supported by The Alfredo Federico Strauss Center for Computational Neuro-imaging | Tel Aviv University
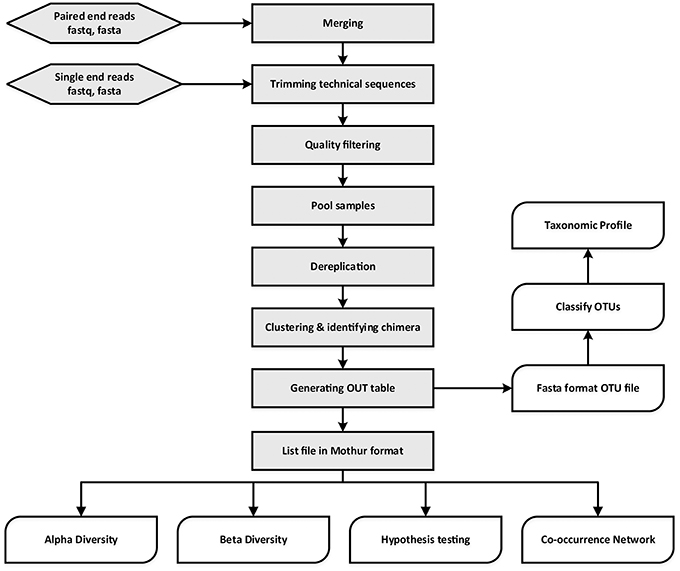
Frontiers | A flexible and economical barcoding approach for highly multiplexed amplicon sequencing of diverse target genes
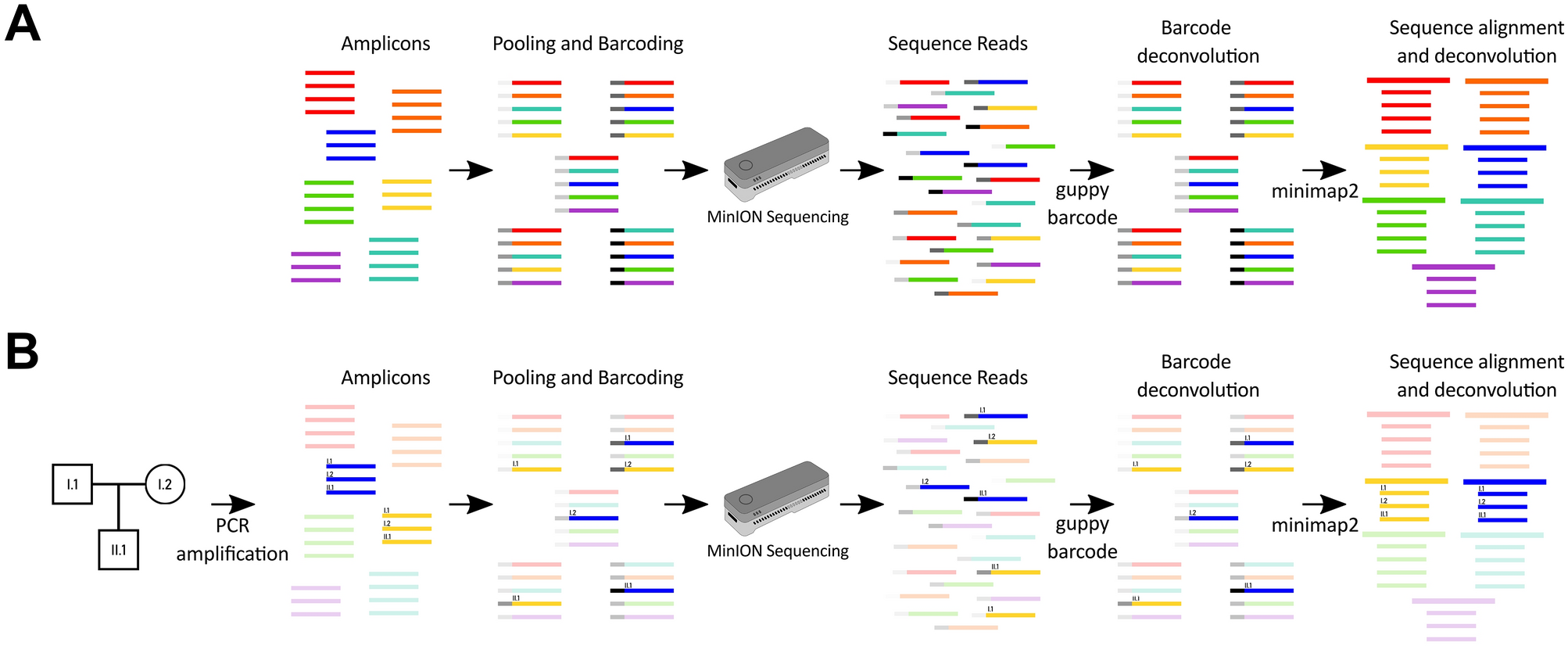
Proof of concept for multiplex amplicon sequencing for mutation identification using the MinION nanopore sequencer | Scientific Reports

Parallel tagged amplicon sequencing of relatively long PCR products using the Illumina HiSeq platform and transcriptome assembly - Feng - 2016 - Molecular Ecology Resources - Wiley Online Library
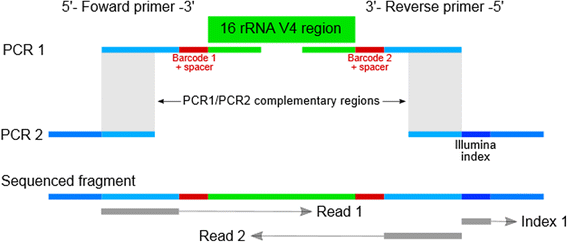
A novel ultra high-throughput 16S rRNA gene amplicon sequencing library preparation method for the Illumina HiSeq platform | Microbiome | Full Text

An optimized, amplicon-based approach for sequencing of SARS-CoV-2 from patient samples using COVIDSeq assay on Illumina MiSeq sequencing platforms - ScienceDirect
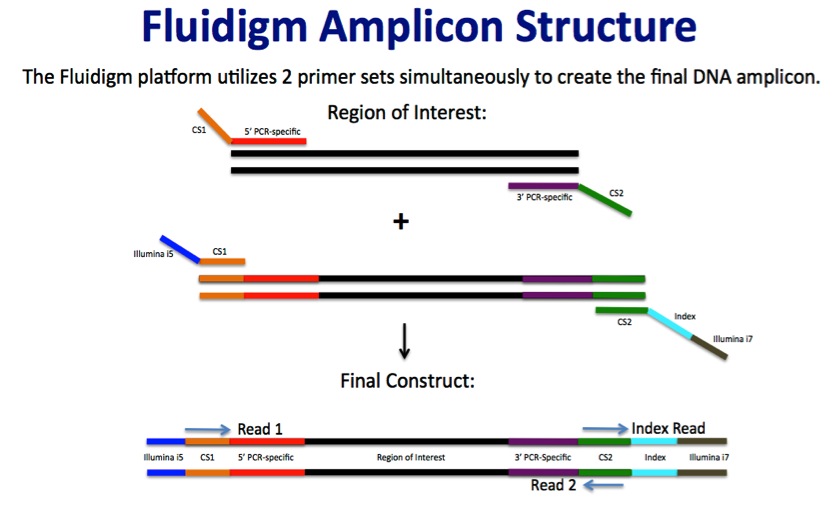
HTDNA: Services/Equipment: Preparation of 16S rRNA (and all other) Amplicons for Illumina Sequencing | Biotech
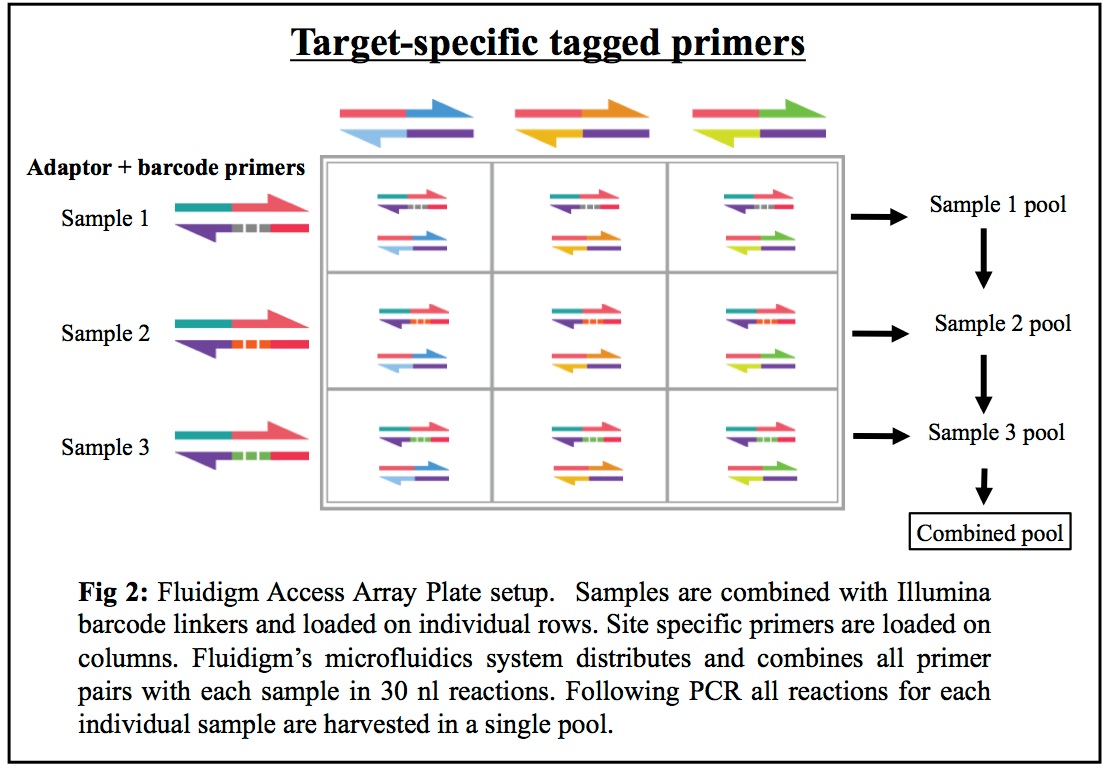
HTDNA: Services/Equipment: Preparation of 16S rRNA (and all other) Amplicons for Illumina Sequencing | Biotech
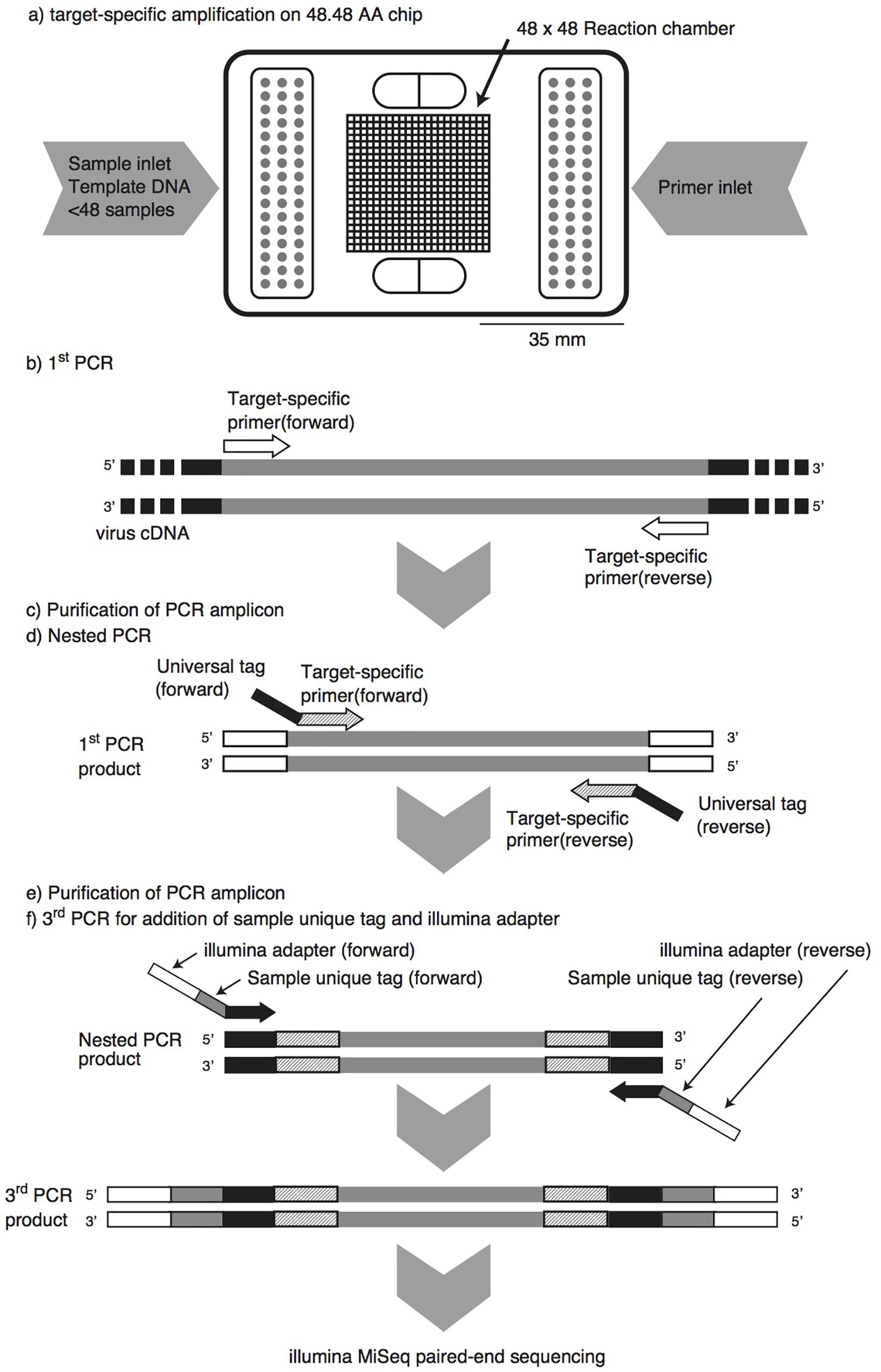
Frontiers | Microfluidic PCR Amplification and MiSeq Amplicon Sequencing Techniques for High-Throughput Detection and Genotyping of Human Pathogenic RNA Viruses in Human Feces, Sewage, and Oysters

Ultrahigh-Throughput Multiplexing and Sequencing of >500-Base-Pair Amplicon Regions on the Illumina HiSeq 2500 Platform | mSystems
Illumina amplicon library preparation through 2-Step PCR amplification. | Download Scientific Diagram
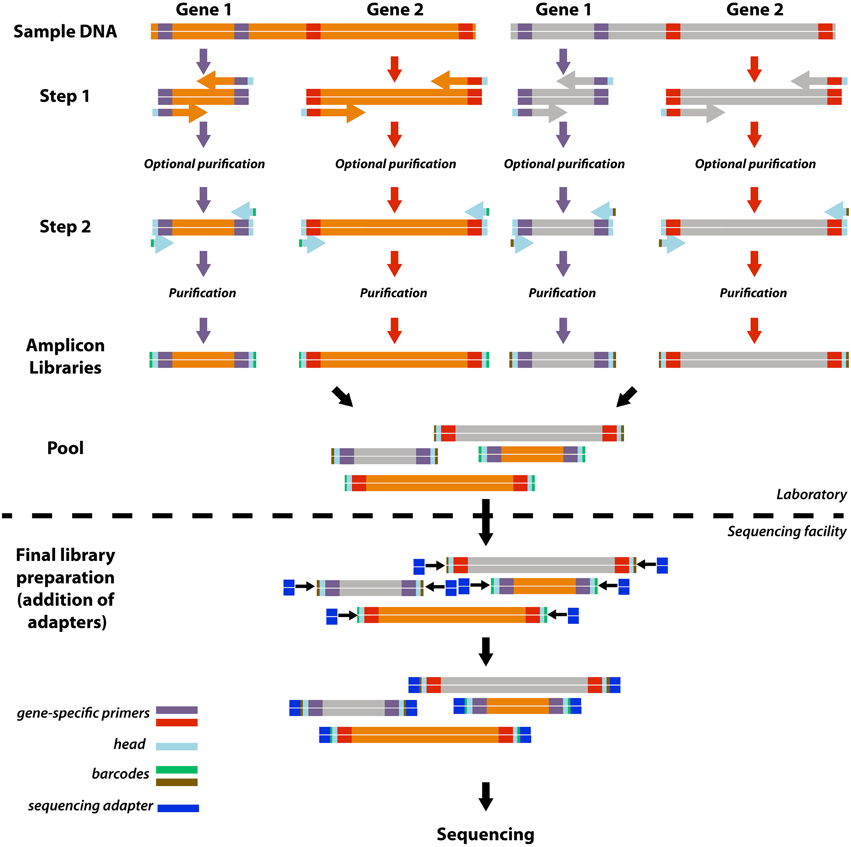
![Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ] Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7786/1/fig-3-2x.jpg)

![Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ] Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7786/1/fig-1-full.png)