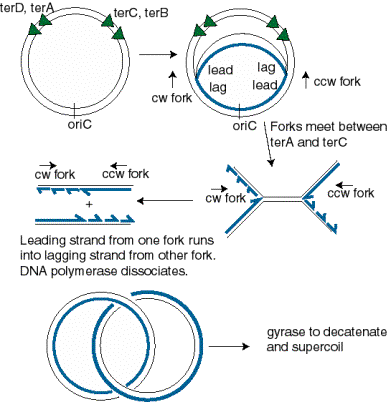
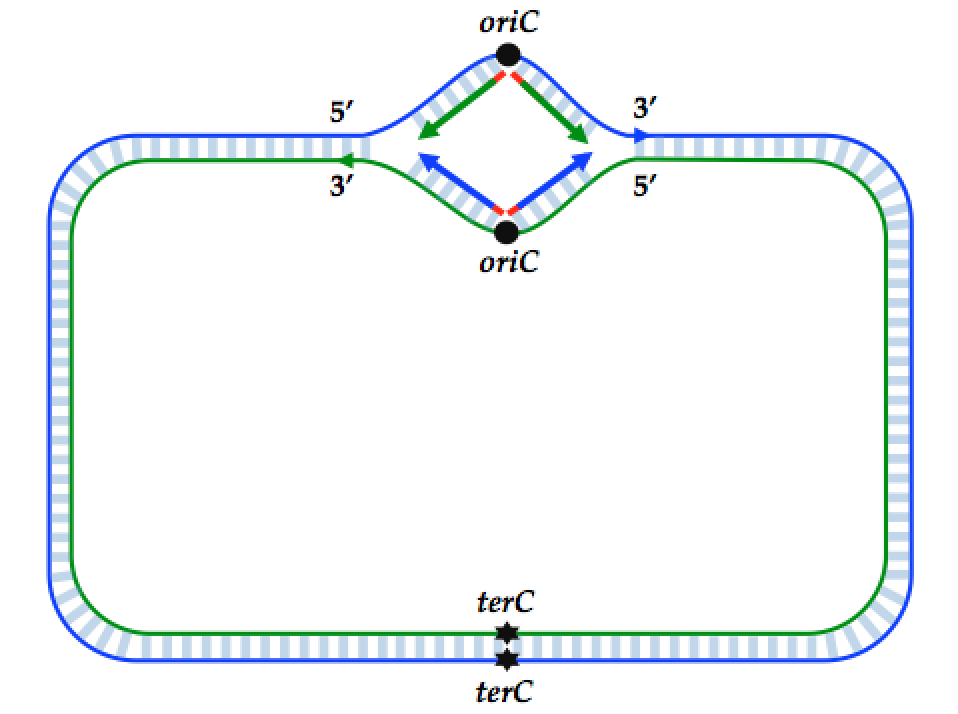
Origin organization and recognition in bacteria. A) Schematic of the... | Download Scientific Diagram

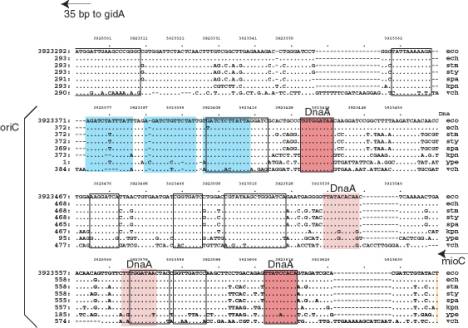
Structural organization of the E. coli oriC region and the sequence of... | Download Scientific Diagram

Differentiation of the DnaA-oriC Subcomplex for DNA Unwinding in a Replication Initiation Complex - ScienceDirect
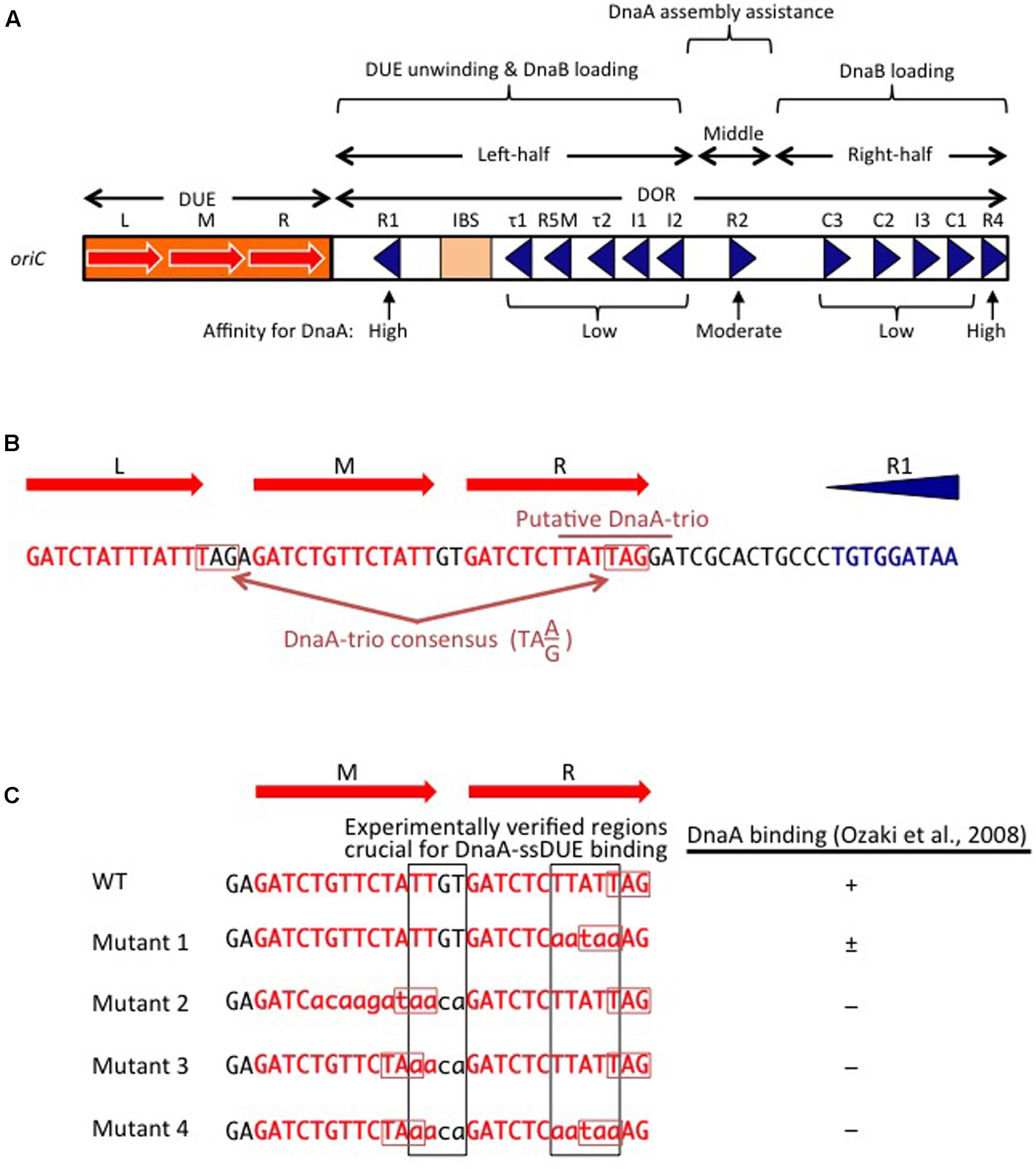
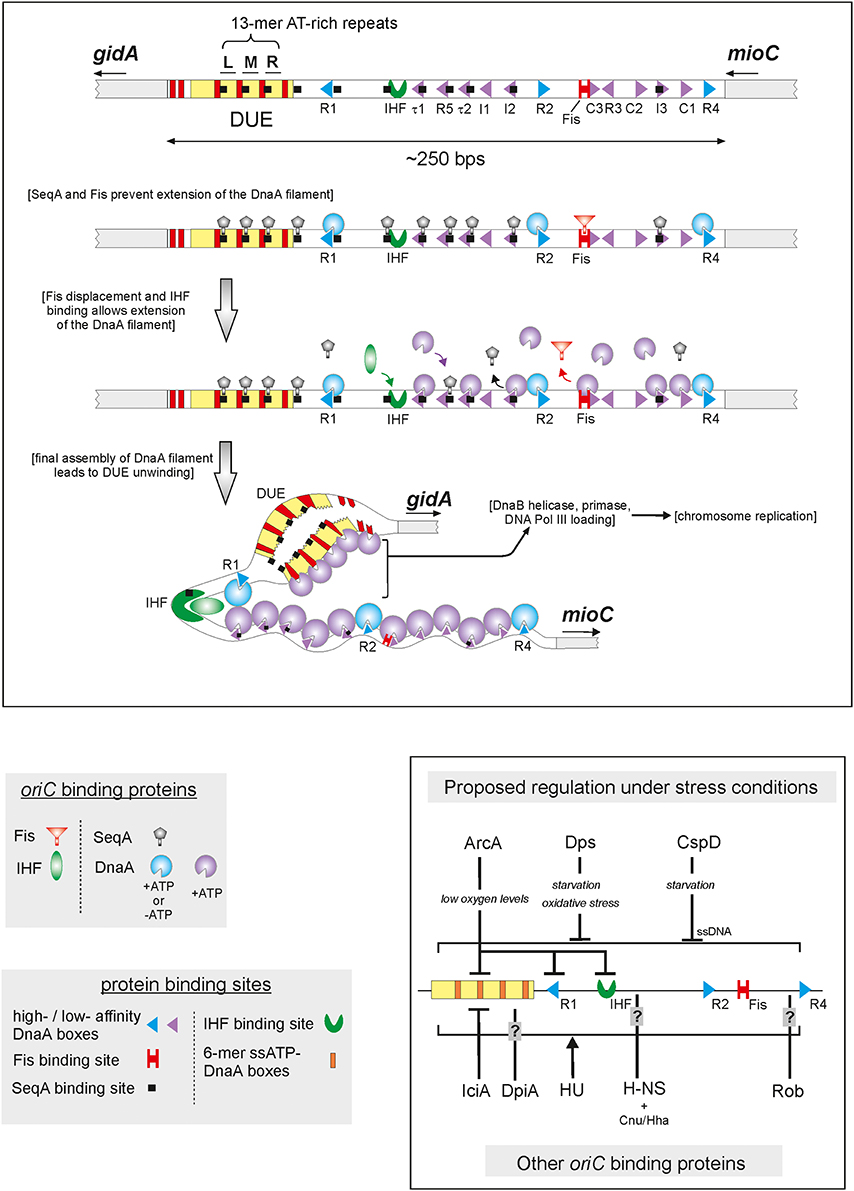
Frontiers | The DnaA Cycle in Escherichia coli: Activation, Function and Inactivation of the Initiator Protein

Structure of oriC, DnaA, the initiation complex, DARS1, and DARS2. (A)... | Download Scientific Diagram

Positions of Fis and DnaA sites at the E.coli oriC shown by sequence... | Download Scientific Diagram
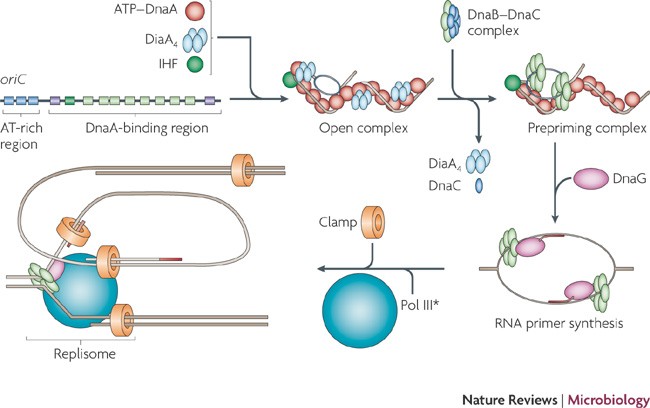
Regulation of the replication cycle: conserved and diverse regulatory systems for DnaA and oriC | Nature Reviews Microbiology

Structures of oriC and DnaA and models for DnaA complexes. A, overall... | Download Scientific Diagram
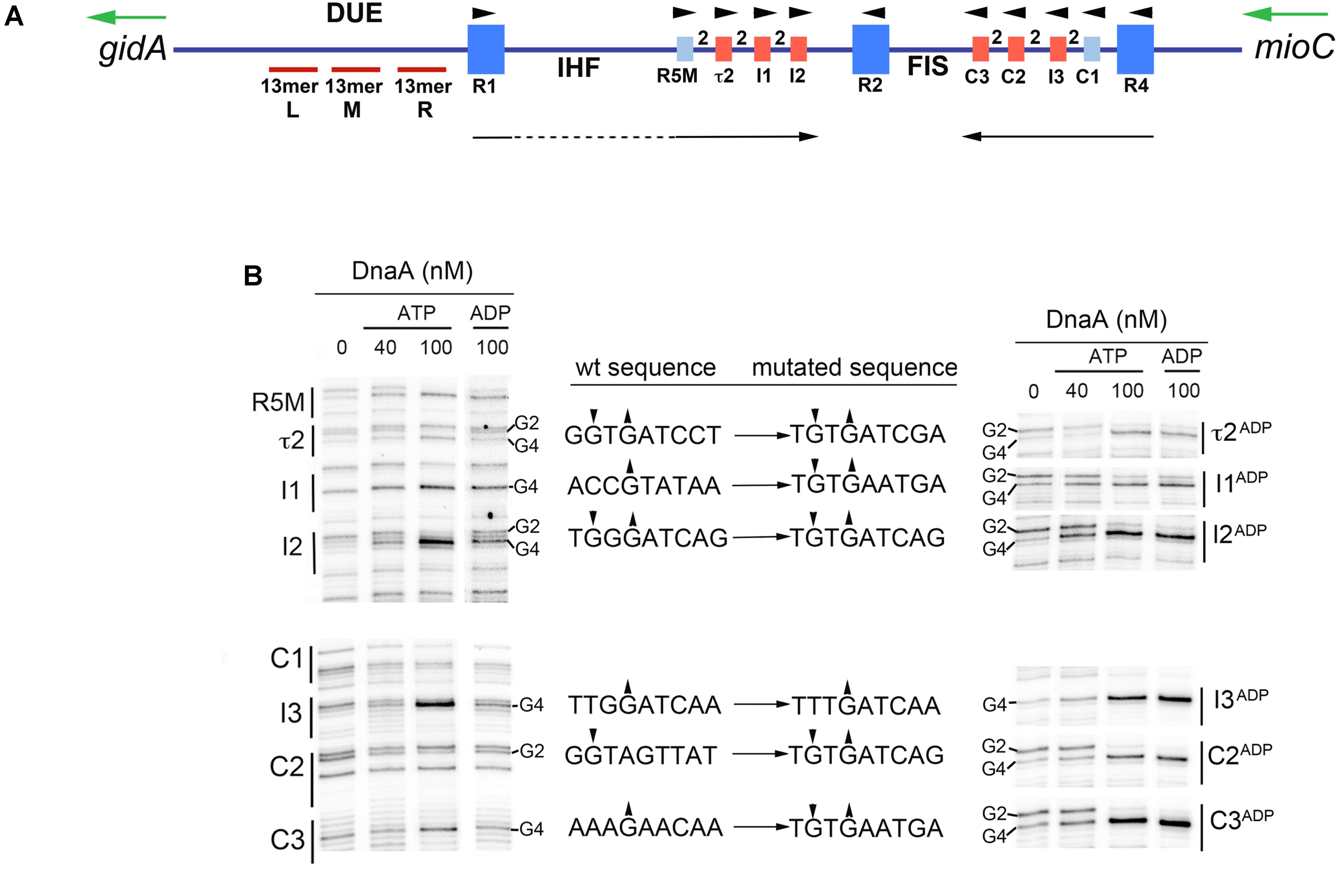
Frontiers | Low Affinity DnaA-ATP Recognition Sites in E. coli oriC Make Non-equivalent and Growth Rate-Dependent Contributions to the Regulated Timing of Chromosome Replication
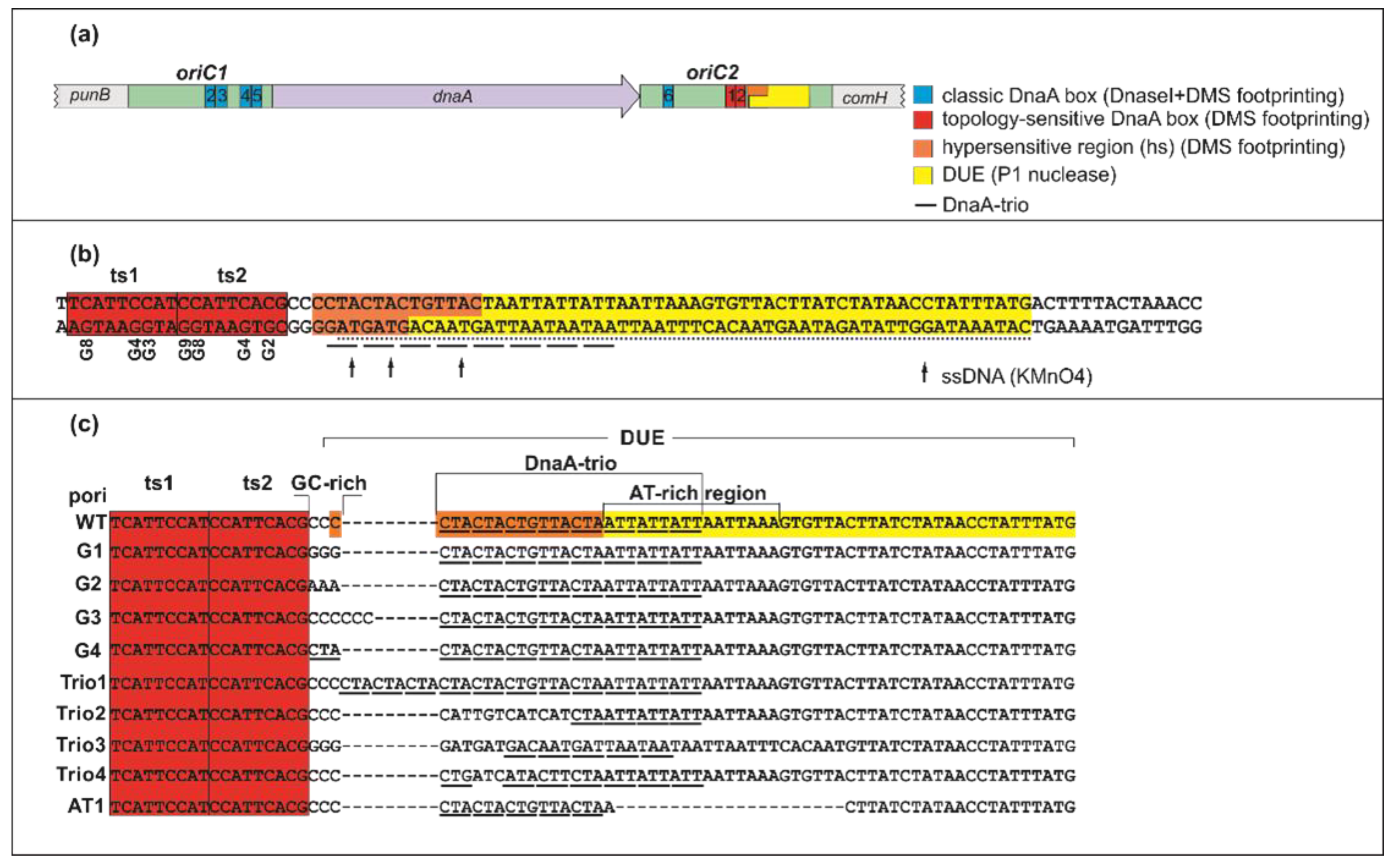
IJMS | Free Full-Text | Putative Cooperative ATP–DnaA Binding to Double-Stranded DnaA Box and Single-Stranded DnaA-Trio Motif upon Helicobacter pylori Replication Initiation Complex Assembly
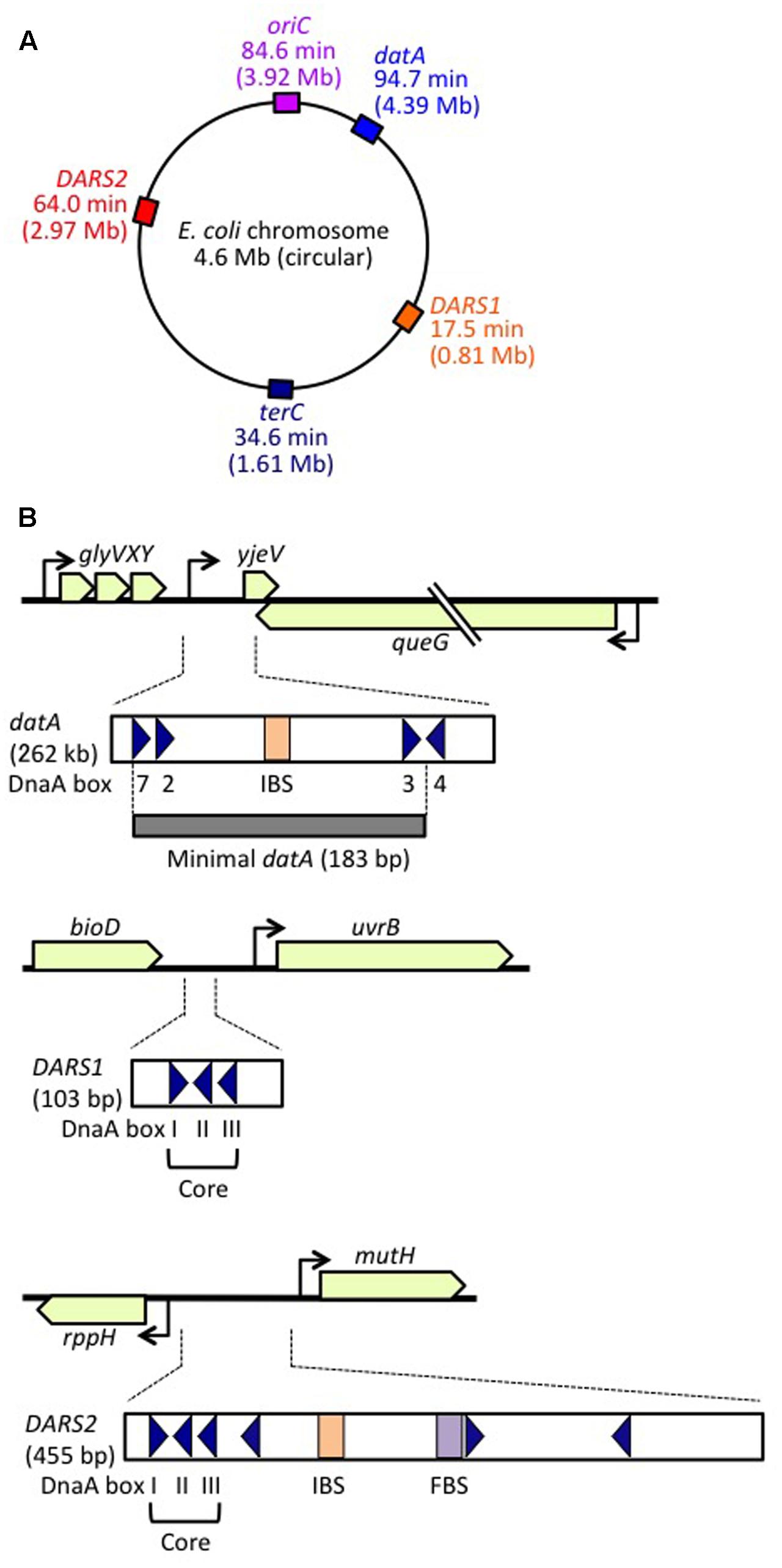
Frontiers | The DnaA Cycle in Escherichia coli: Activation, Function and Inactivation of the Initiator Protein

Near-atomic structural model for bacterial DNA replication initiation complex and its functional insights | PNAS

Positioning and the specific sequence of each 13‐mer motif are critical for activity of the plasmid RK2 replication origin - Kowalczyk - 2005 - Molecular Microbiology - Wiley Online Library