6 (A) Sequence alignment of template CYP120A1 (PDB ID 2VE3) and amino... | Download Scientific Diagram
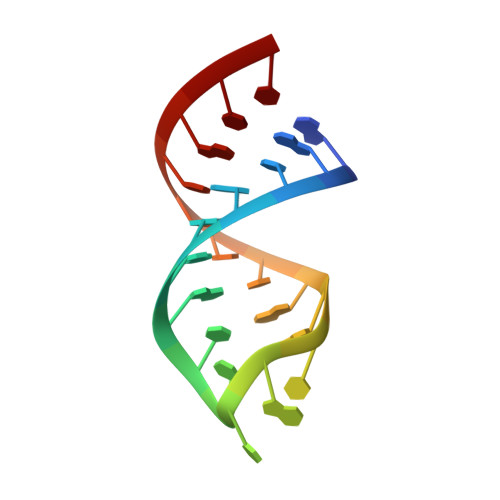
RCSB PDB - 1SZY: Solution structure of ITALY1 ("Initiator tRNA Anticodon Loop from Yeast"), an unmodified 21-nt RNA with the sequence of the anticodon stem-loop of yeast initiator tRNA

RCSB PDB - 140D: SOLUTION STRUCTURE OF A CONSERVED DNA SEQUENCE FROM THE HIV-1 GENOME: RESTRAINED MOLECULAR DYNAMICS SIMULATION WITH DISTANCE AND TORSION ANGLE RESTRAINTS DERIVED FROM TWO-DIMENSIONAL NMR SPECTRA

Figure S6: Sequence chain view of the structure of DyP4 F194Y variant... | Download Scientific Diagram

RCSB PDB - 1XE8: Crystal structure of the YML079w protein from Saccharomyces cerevisiae reveals a new sequence family of the jelly roll fold.
Sequence display for secondary structure entities in PDB model 3GUU... | Download Scientific Diagram

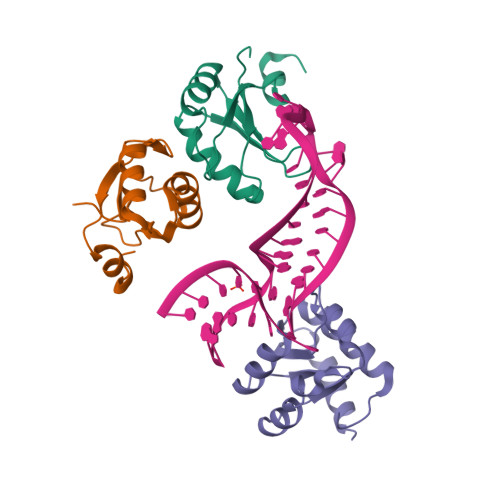
RCSB PDB - 6PAX: CRYSTAL STRUCTURE OF THE HUMAN PAX-6 PAIRED DOMAIN-DNA COMPLEX REVEALS A GENERAL MODEL FOR PAX PROTEIN-DNA INTERACTIONS

RCSB PDB - 1A8C: PRIMARY SEQUENCE AND SOLUTION CONFORMATION OF FERROCYTOCHROME C-552 FROM NITROSOMONAS EUROPAEA, NMR, MEAN STRUCTURE REFINED WITHOUT HYDROGEN BOND CONSTRAINTS

RCSB PDB - 1ND1: Amino acid sequence and crystal structure of BaP1, a metalloproteinase from Bothrops asper snake venom that exerts multiple tissue-damaging activities.