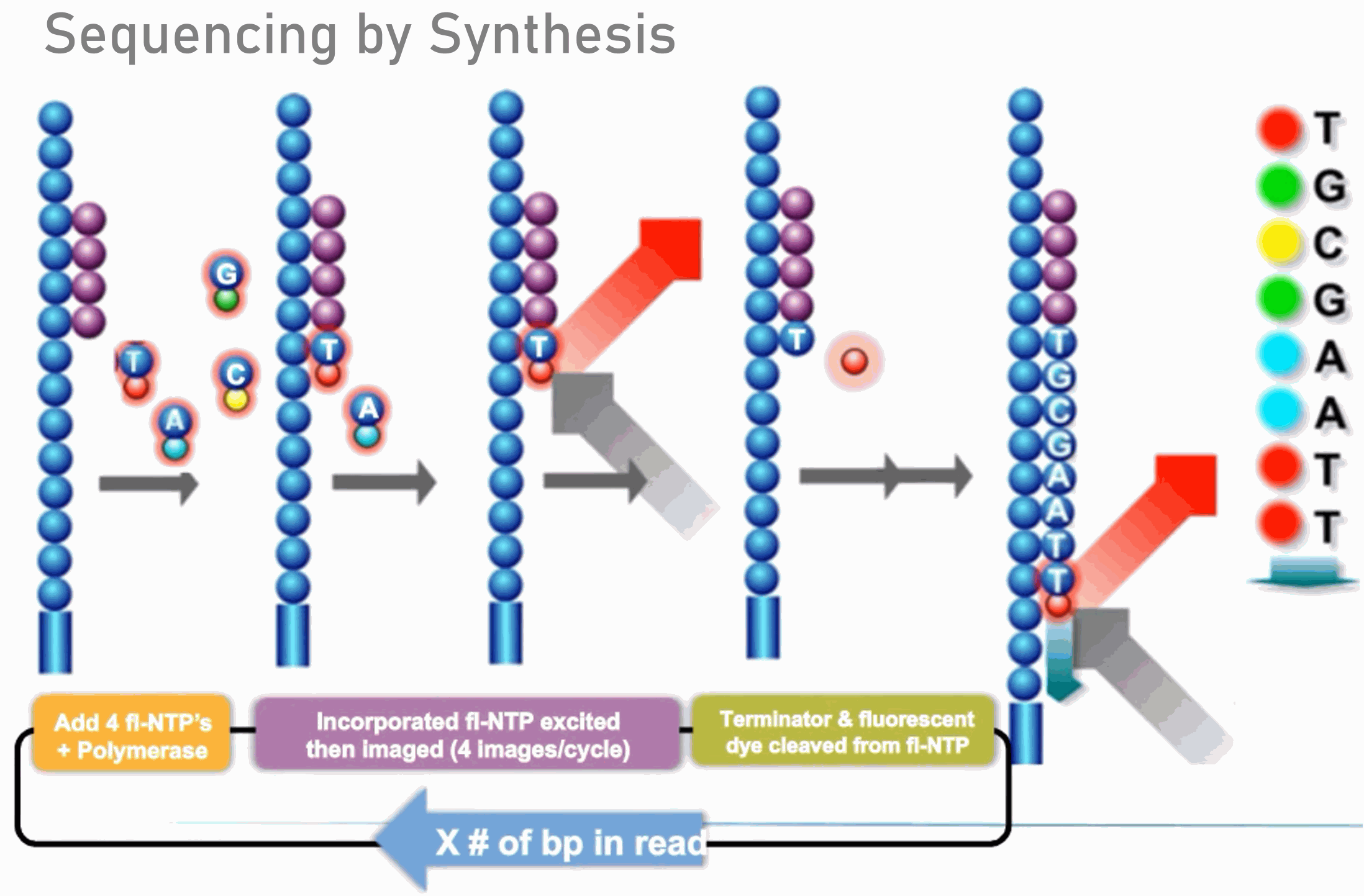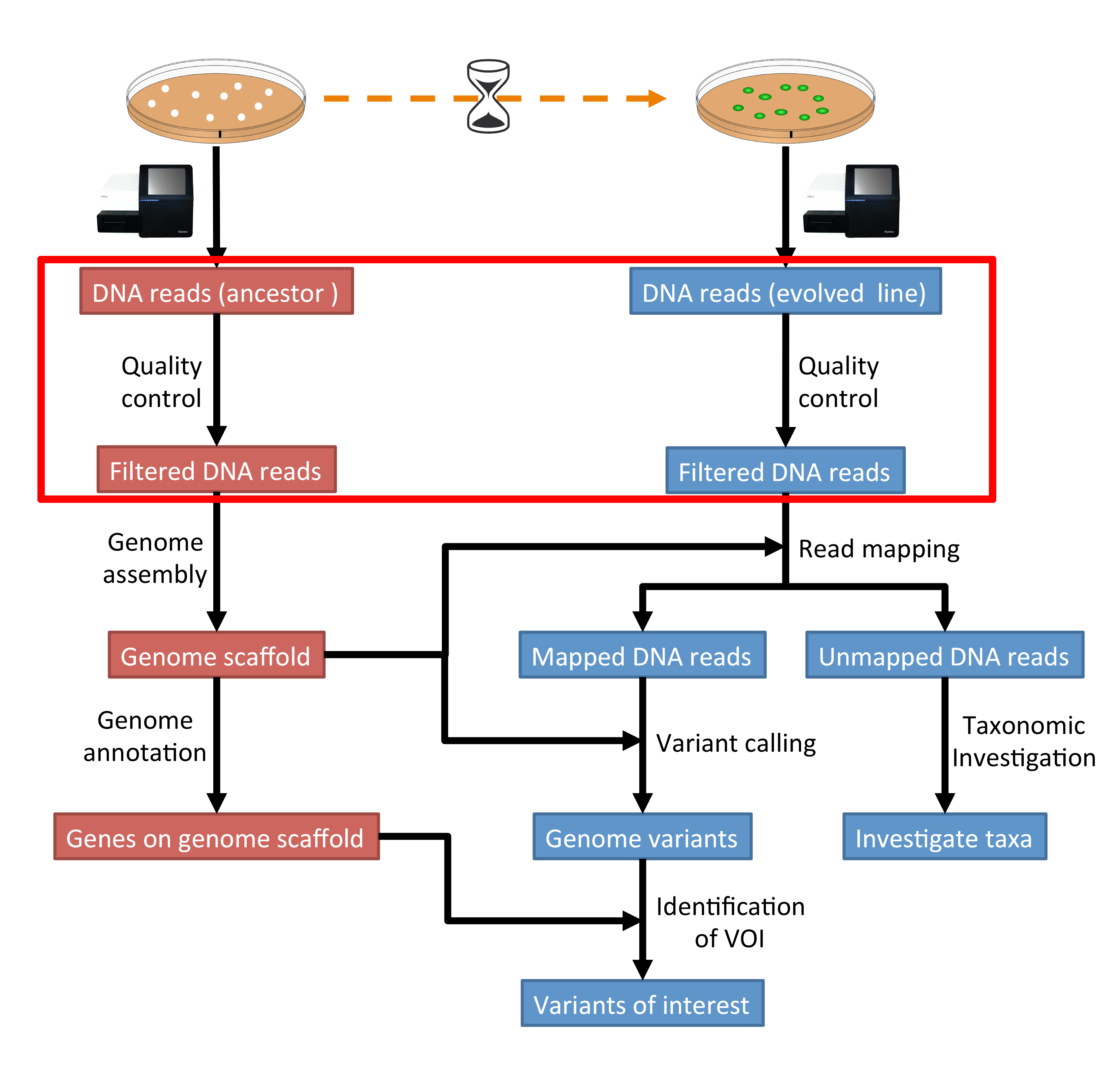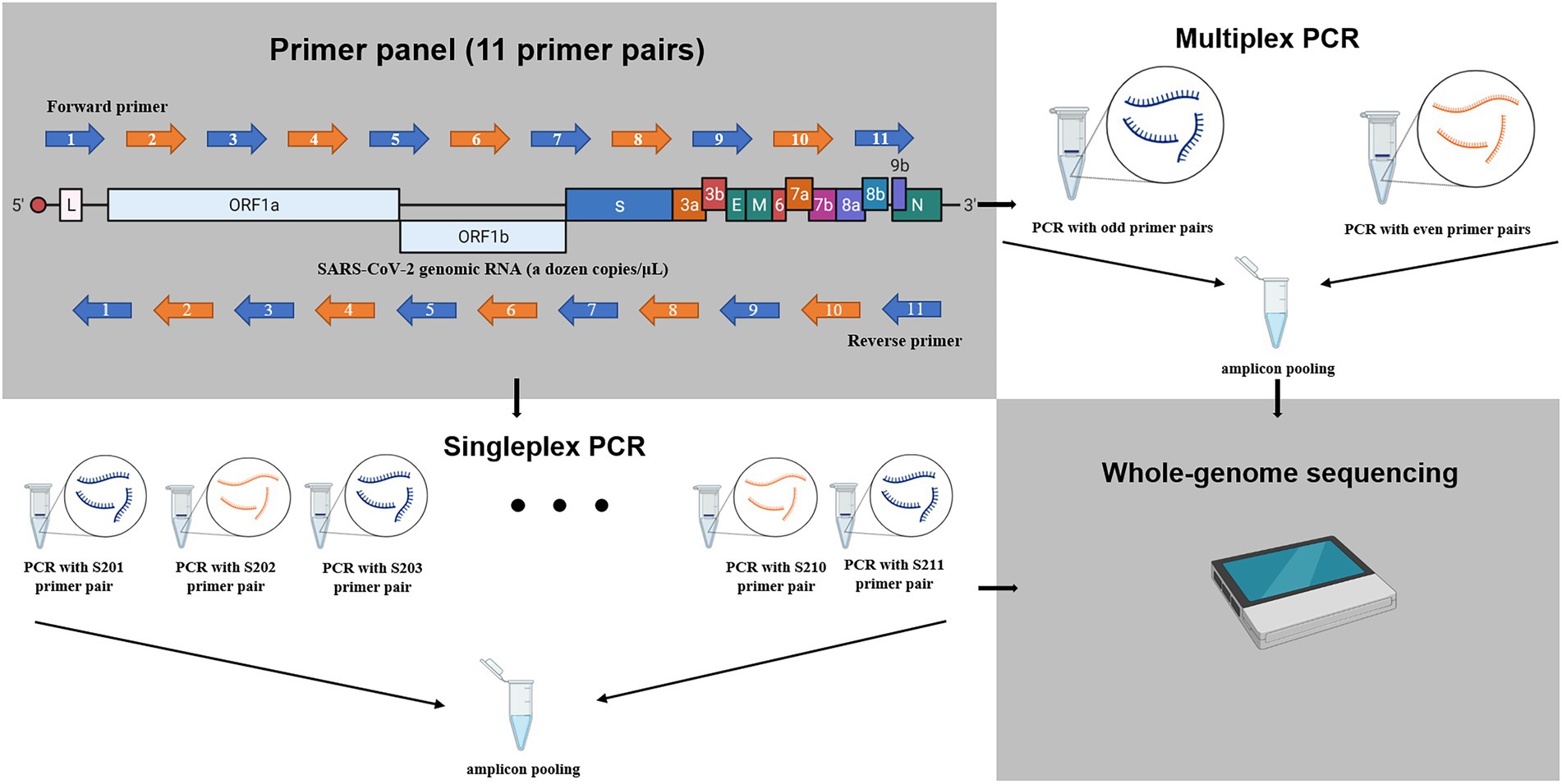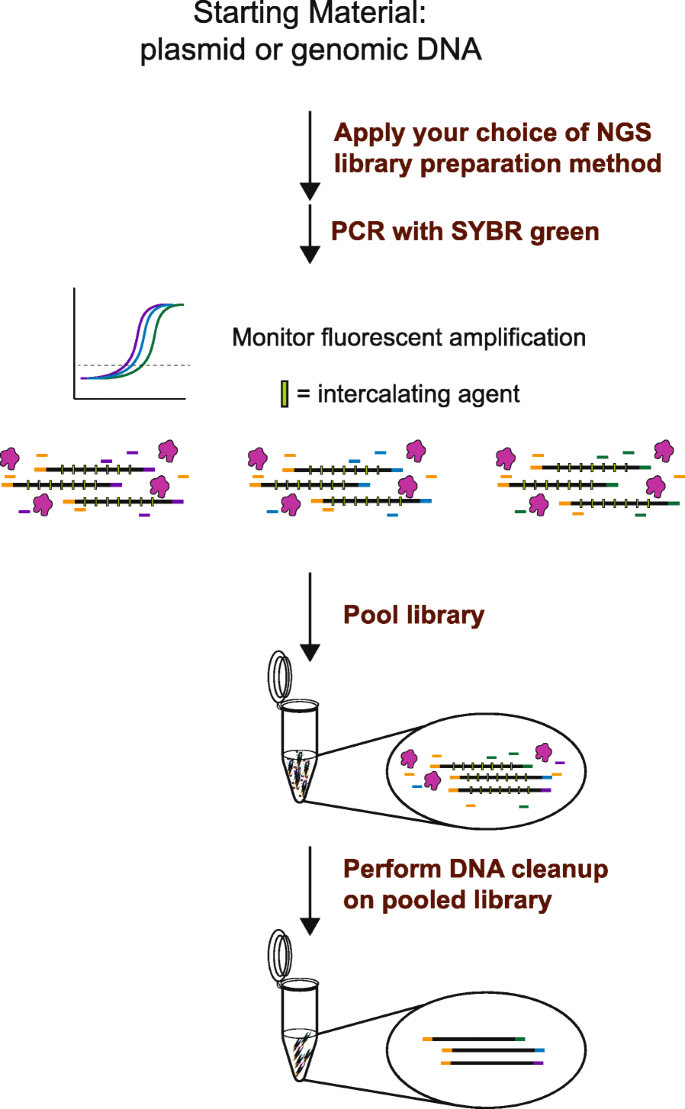
Fluorescent amplification for next generation sequencing (FA-NGS) library preparation | BMC Genomics | Full Text
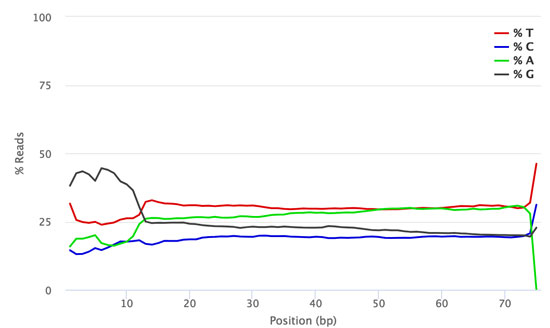
Strategies for Achieving High Sequencing Accuracy for Low Diversity Samples and Avoiding Sample Bleeding Using Illumina Platform | PLOS ONE
Quality scores of sequencing runs. (A) Mean sequencing quality scores... | Download Scientific Diagram

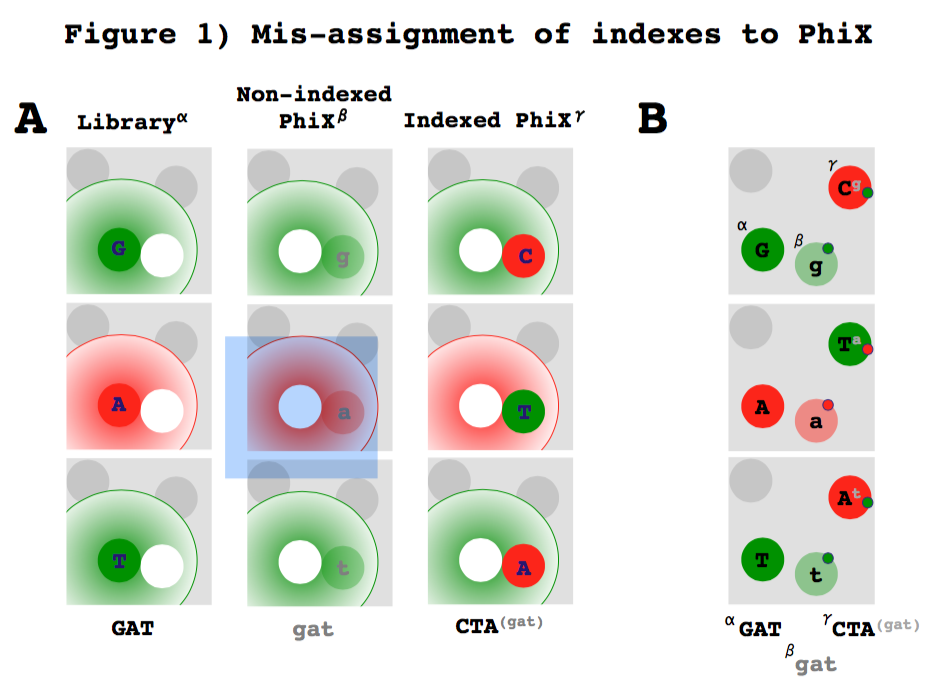
How much PhiX spike in is recommended when sequencing low diversity libraries on Illumina platforms? - Illumina Knowledge
Anweisung zum Erstellen des Samplesheets für einen PhiX-Validierungslauf auf dem MiSeq-System mit Illumina Experiment Manager