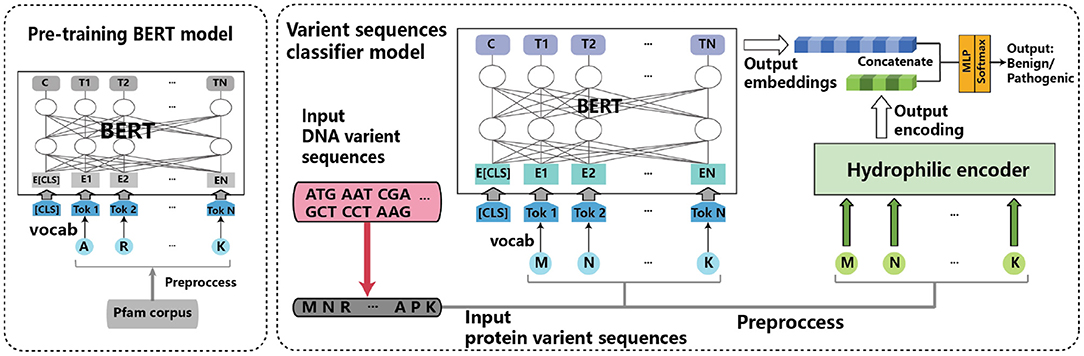
Structure-aware protein solubility prediction from sequence through graph convolutional network and predicted contact map | Journal of Cheminformatics | Full Text

The 4-step flowchart demonstrating our method for using word embedding... | Download Scientific Diagram
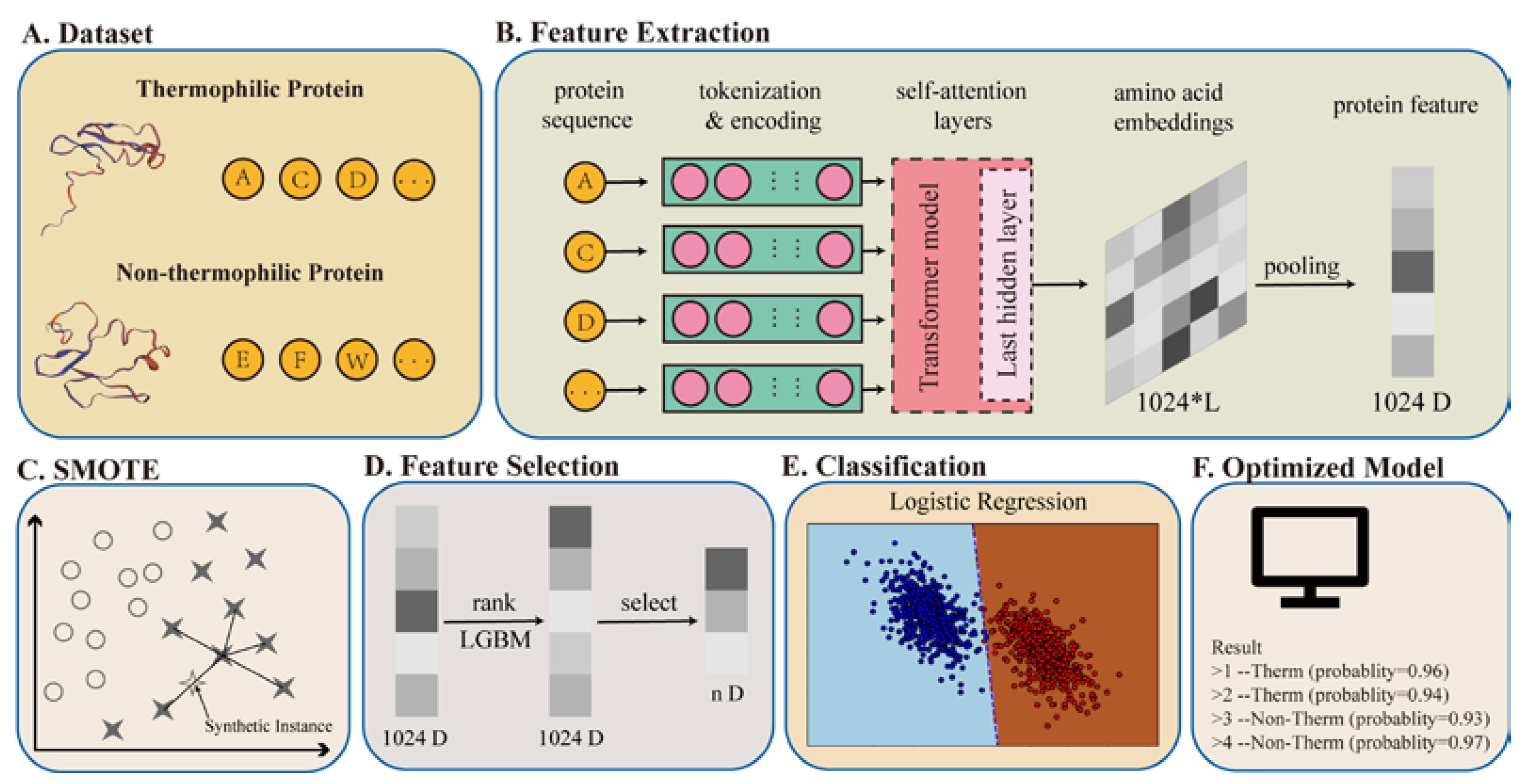
Applied Sciences | Free Full-Text | Identification of Thermophilic Proteins Based on Sequence-Based Bidirectional Representations from Transformer- Embedding Features

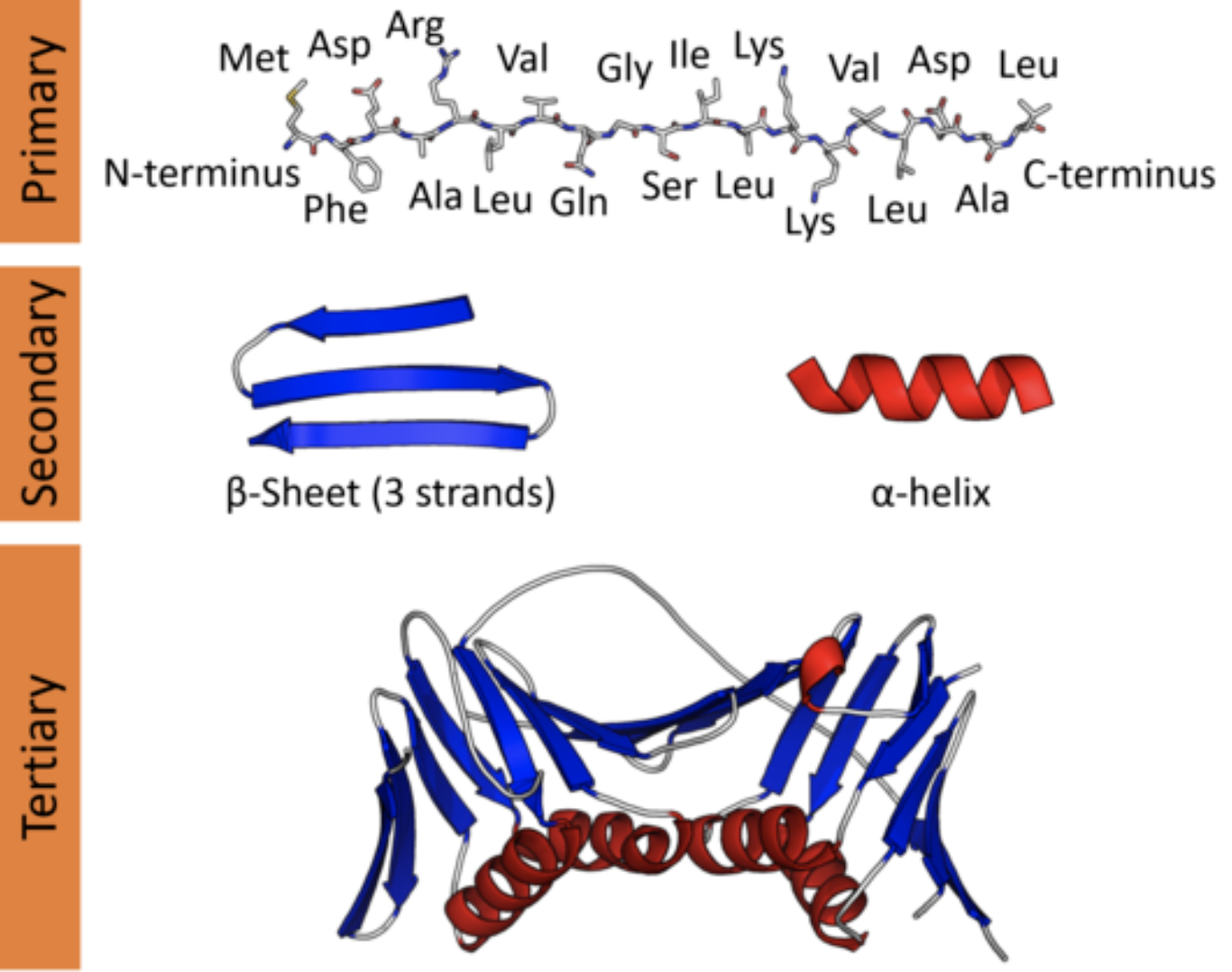
Artificial intelligence method to design and fold alpha-helical structural proteins from the primary amino acid sequence | bioRxiv
![An integration of deep learning with feature embedding for protein–protein interaction prediction [PeerJ] An integration of deep learning with feature embedding for protein–protein interaction prediction [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7126/1/fig-3-2x.jpg)
An integration of deep learning with feature embedding for protein–protein interaction prediction [PeerJ]

Deep hierarchical embedding for simultaneous modeling of GPCR proteins in a unified metric space | Scientific Reports
Sequence embedding method in PhaTYP. The block in "Protein Sentence"... | Download Scientific Diagram

Conjoint Feature Representation of GO and Protein Sequence for PPI Prediction Based on an Inception RNN Attention Network - ScienceDirect
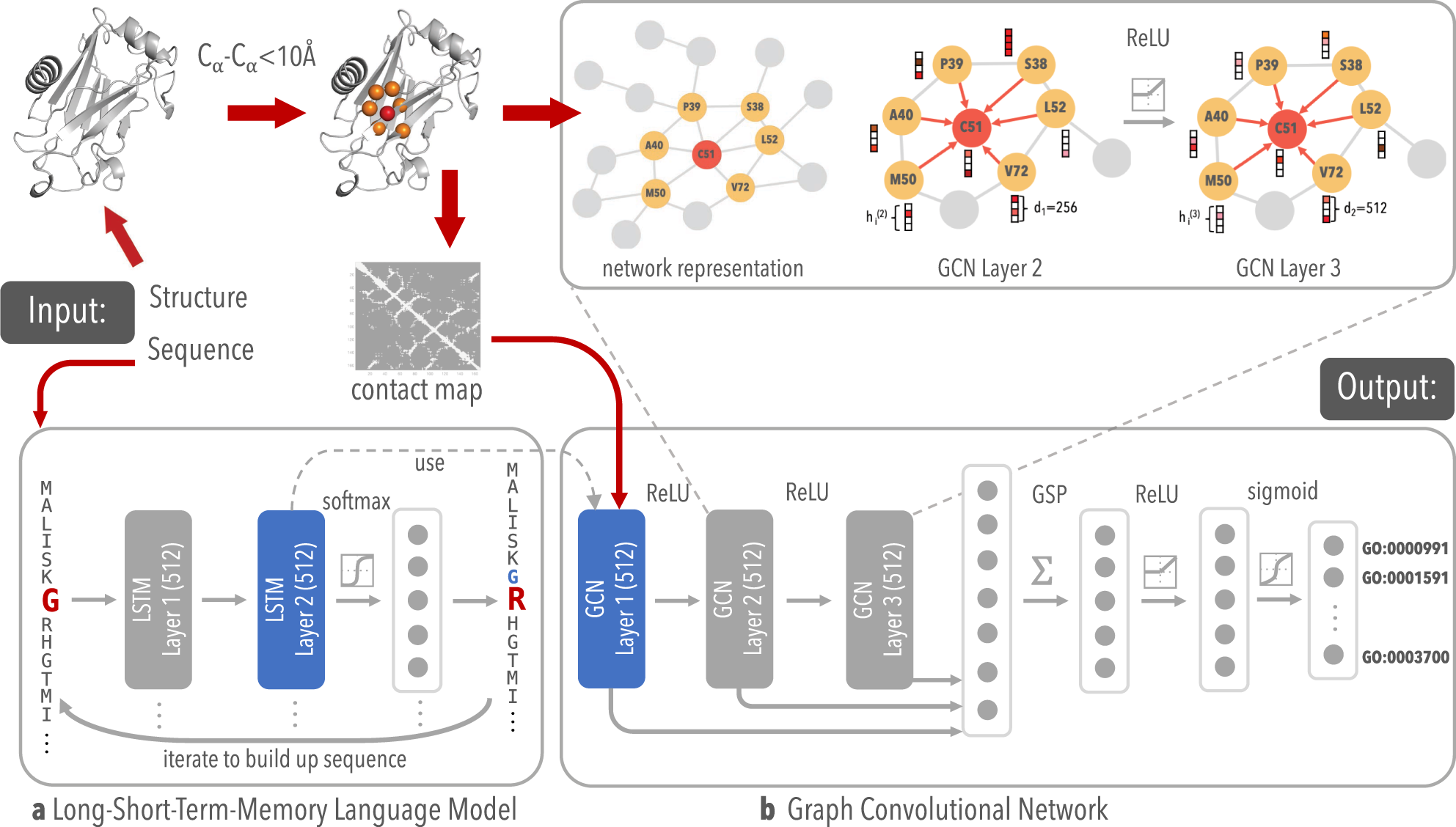
Structure-based protein function prediction using graph convolutional networks | Nature Communications
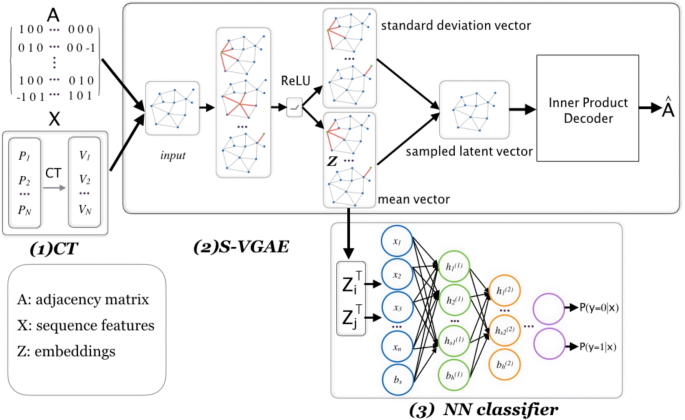
Graph-based prediction of Protein-protein interactions with attributed signed graph embedding | BMC Bioinformatics | Full Text

Learning the molecular grammar of protein condensates from sequence determinants and embeddings | PNAS



![PDF] Learning protein sequence embeddings using information from structure | Semantic Scholar PDF] Learning protein sequence embeddings using information from structure | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/3637fcccc758786ae1c6529ab22fe85ea98e9c36/2-Figure1-1.png)
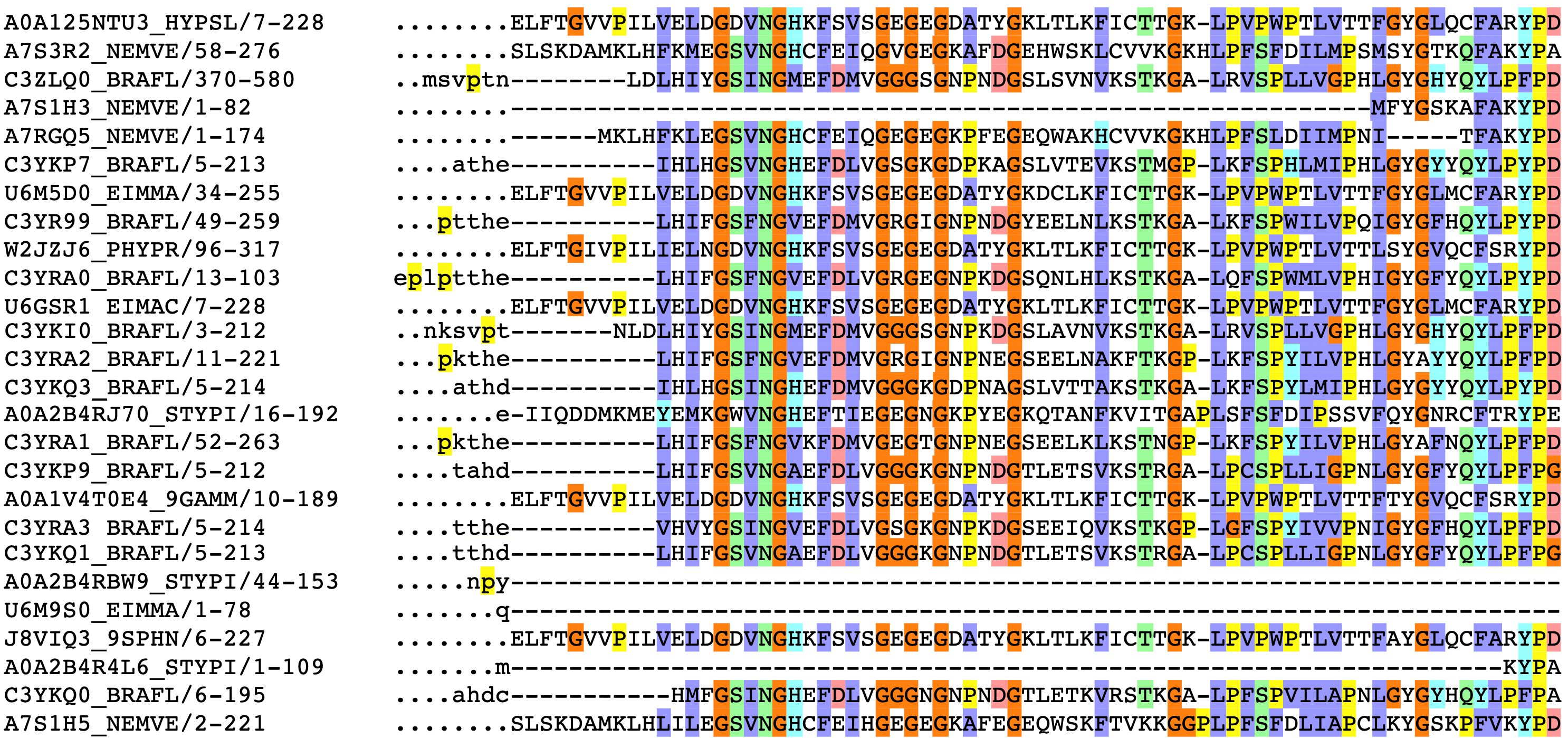
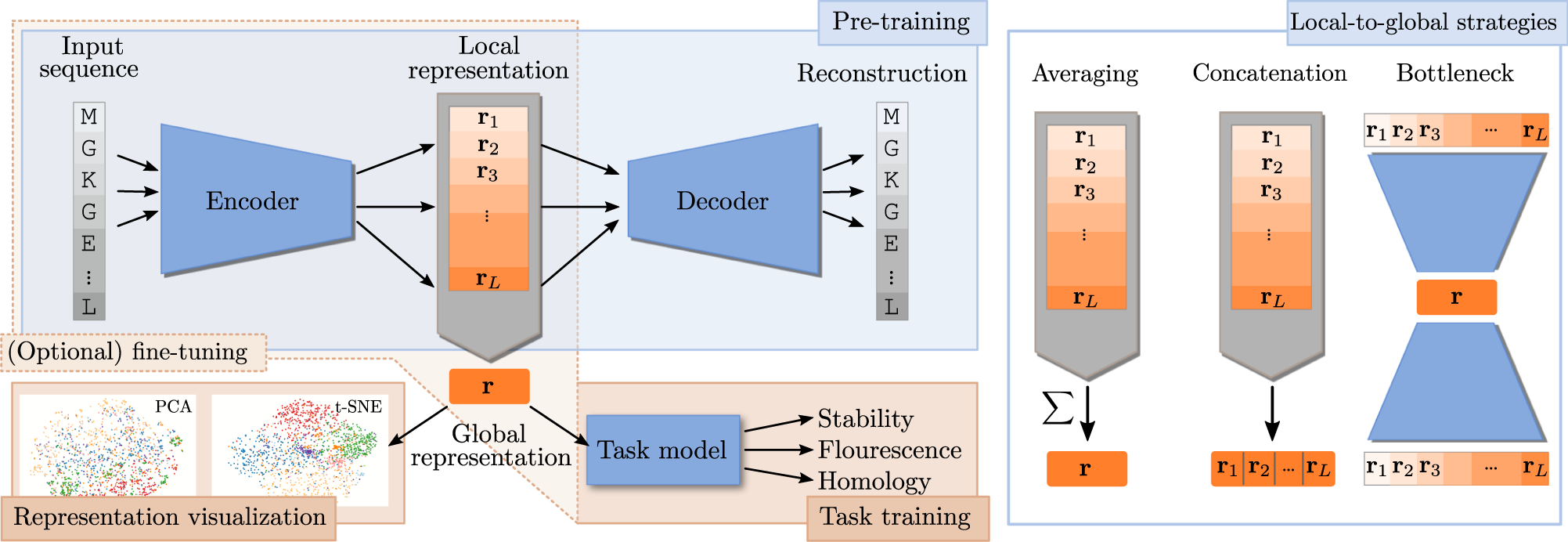
![PDF] Learning protein sequence embeddings using information from structure | Semantic Scholar PDF] Learning protein sequence embeddings using information from structure | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/3637fcccc758786ae1c6529ab22fe85ea98e9c36/13-Figure2-1.png)