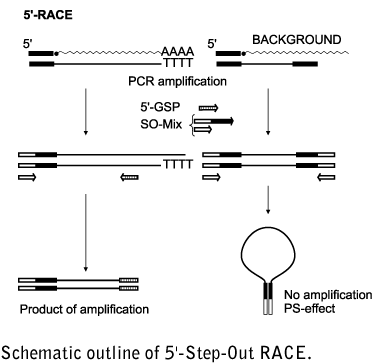
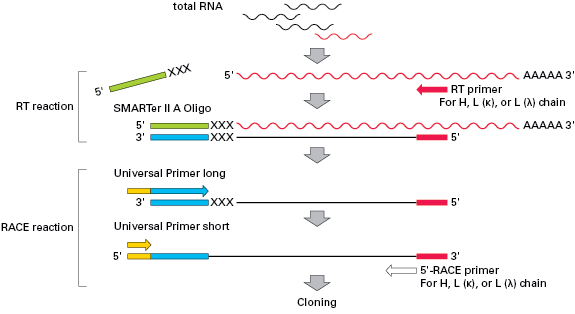
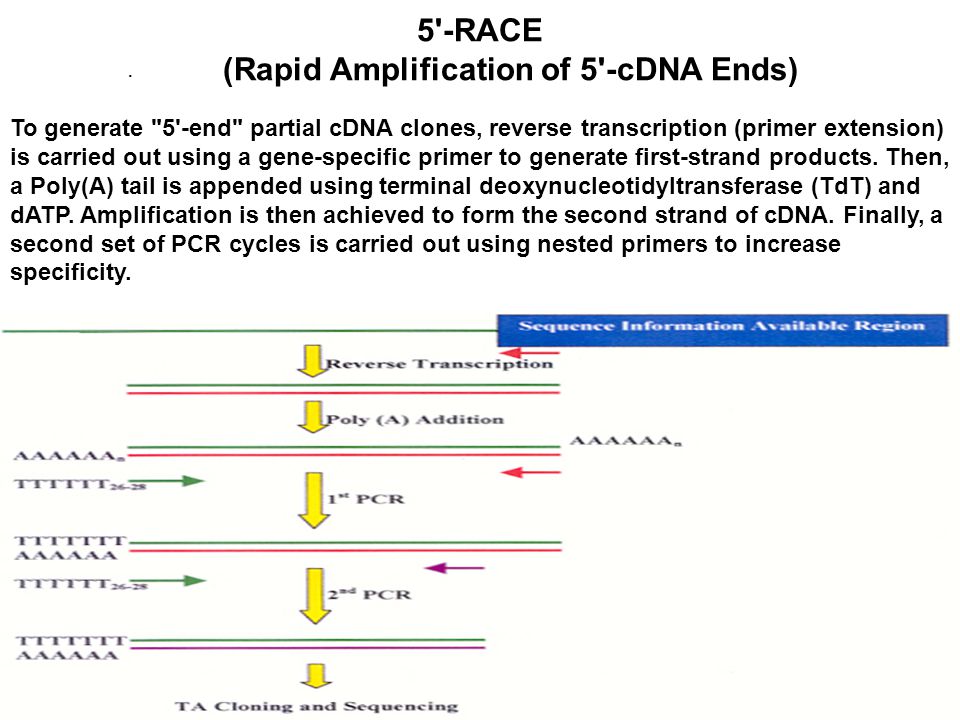
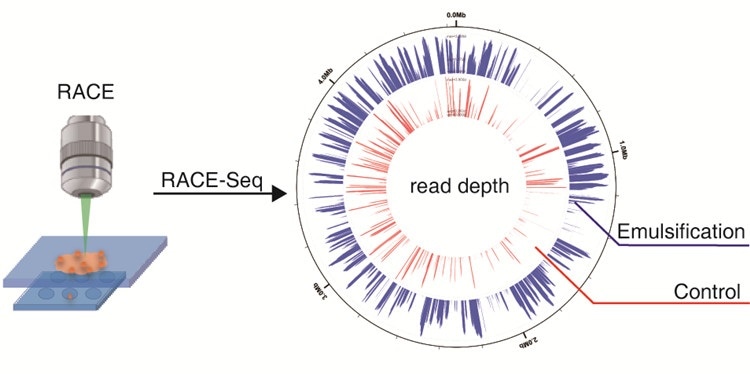
Schematic representation of the 5′-RACE PCR procedure optimized for the... | Download Scientific Diagram

A versatile 5′ RACE-Seq methodology for the accurate identification of the 5′ termini of mRNAs | BMC Genomics | Full Text

miR-RACE – An Approach to Determine the Sequence of Computationally Identified miRNAs | RNA-Seq Blog

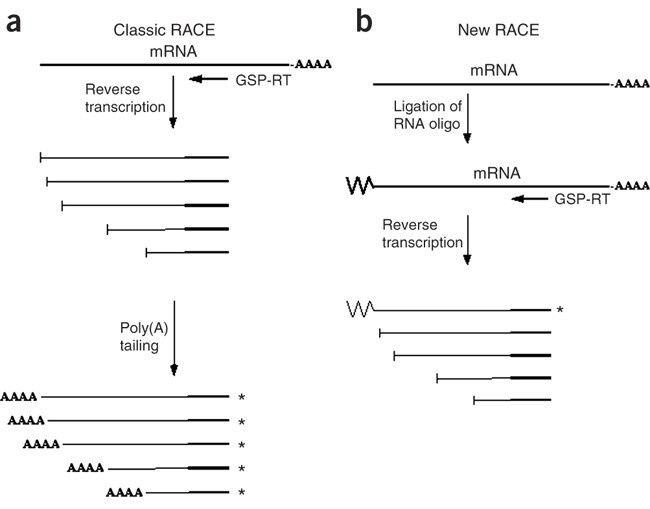
3′ RACE with deep sequencing, schematic of method used. a Ligation of... | Download Scientific Diagram
![PDF] Using Next-Generation Sequencing to Explore Genetics and Race in the High School Classroom | Semantic Scholar PDF] Using Next-Generation Sequencing to Explore Genetics and Race in the High School Classroom | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/4b8debef7cdd5efe13076b647f5b7cee7917f54b/3-Figure1-1.png)
PDF] Using Next-Generation Sequencing to Explore Genetics and Race in the High School Classroom | Semantic Scholar
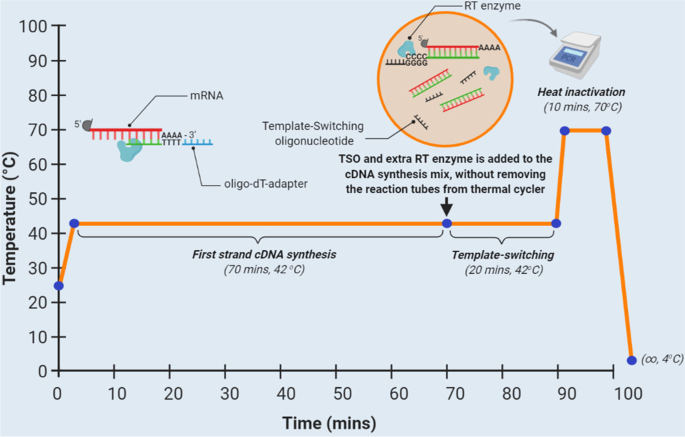
A versatile 5′ RACE-Seq methodology for the accurate identification of the 5′ termini of mRNAs | BMC Genomics | Full Text

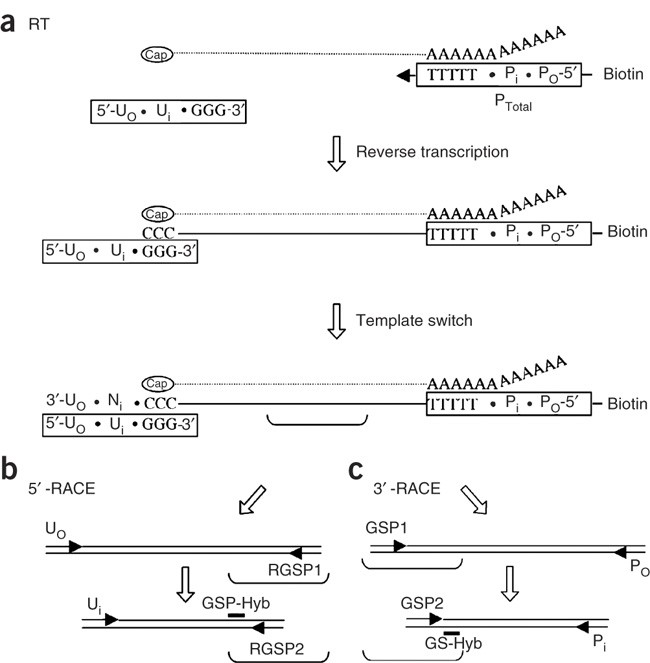
a. Determination of the precise miRNA sequence by 5′ and 3′ miR-RACE.... | Download Scientific Diagram

RACE-SEQ and Population-Wide Polymorphism Susceptibility Testing for Endonucleolytically Active, RNA-Targeting Therapeutics | SpringerLink
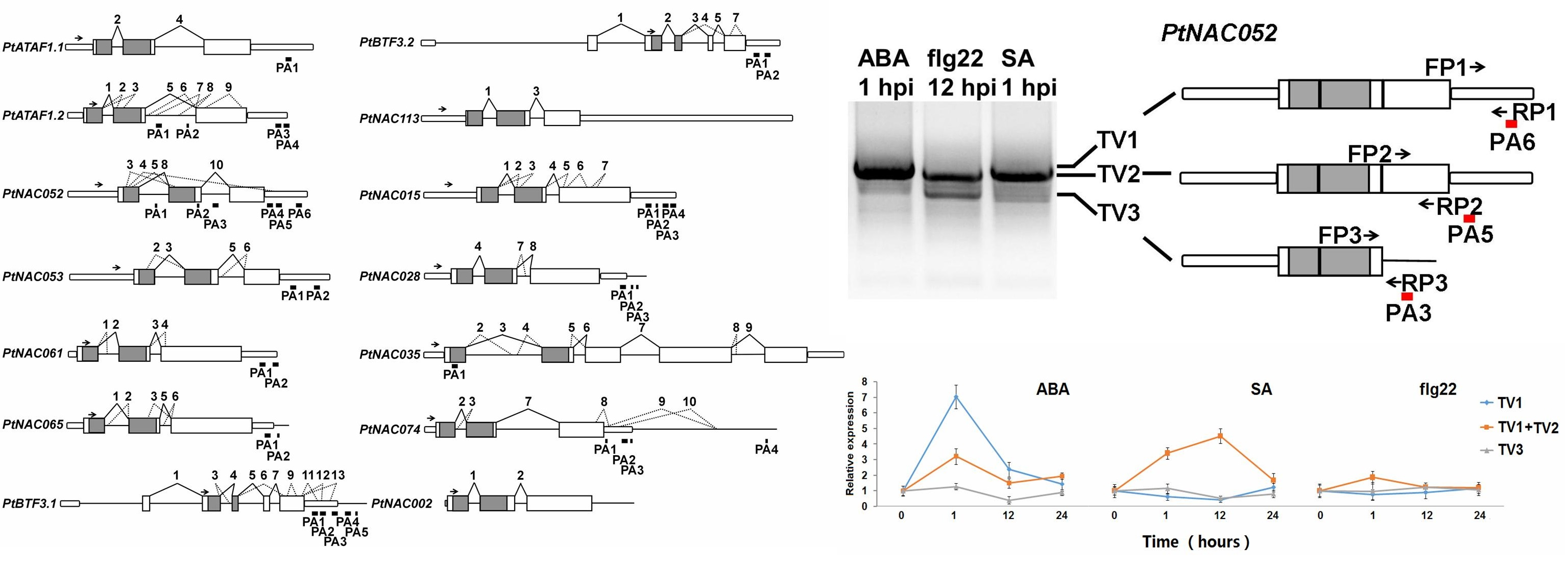
Molecules | Free Full-Text | Capturing the Alternative Cleavage and Polyadenylation Sites of 14 NAC Genes in Populus Using a Combination of 3′- RACE and High-Throughput Sequencing