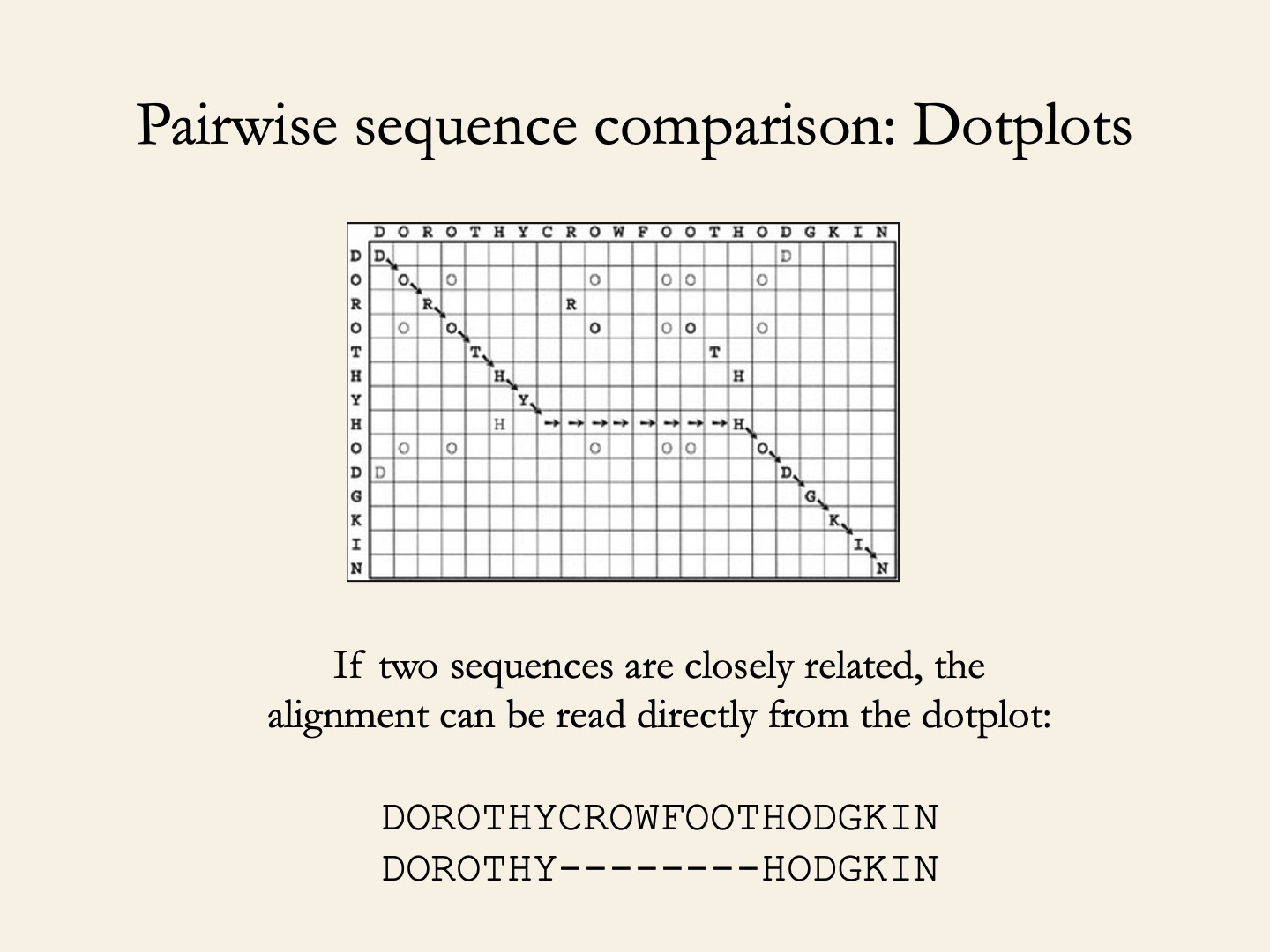
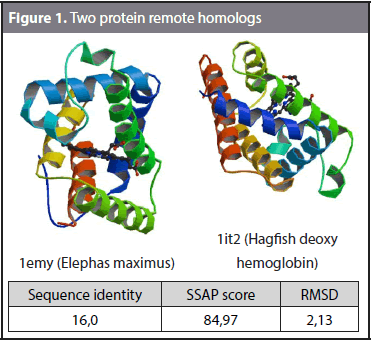
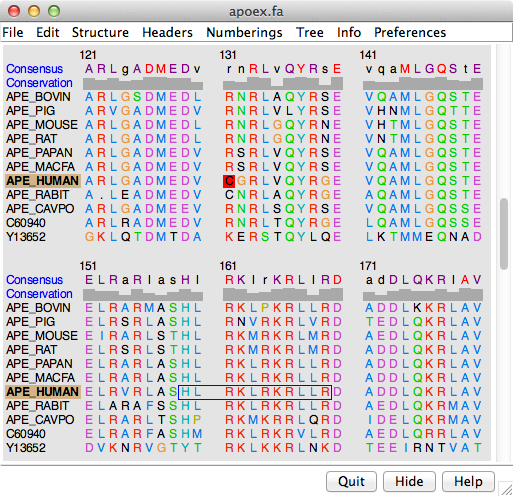
Amino acid identity table and sequence alignments of Act d 10, Ara h 9,... | Download Scientific Diagram

Difference Between Similarity and Identity in Sequence Alignment | Compare the Difference Between Similar Terms
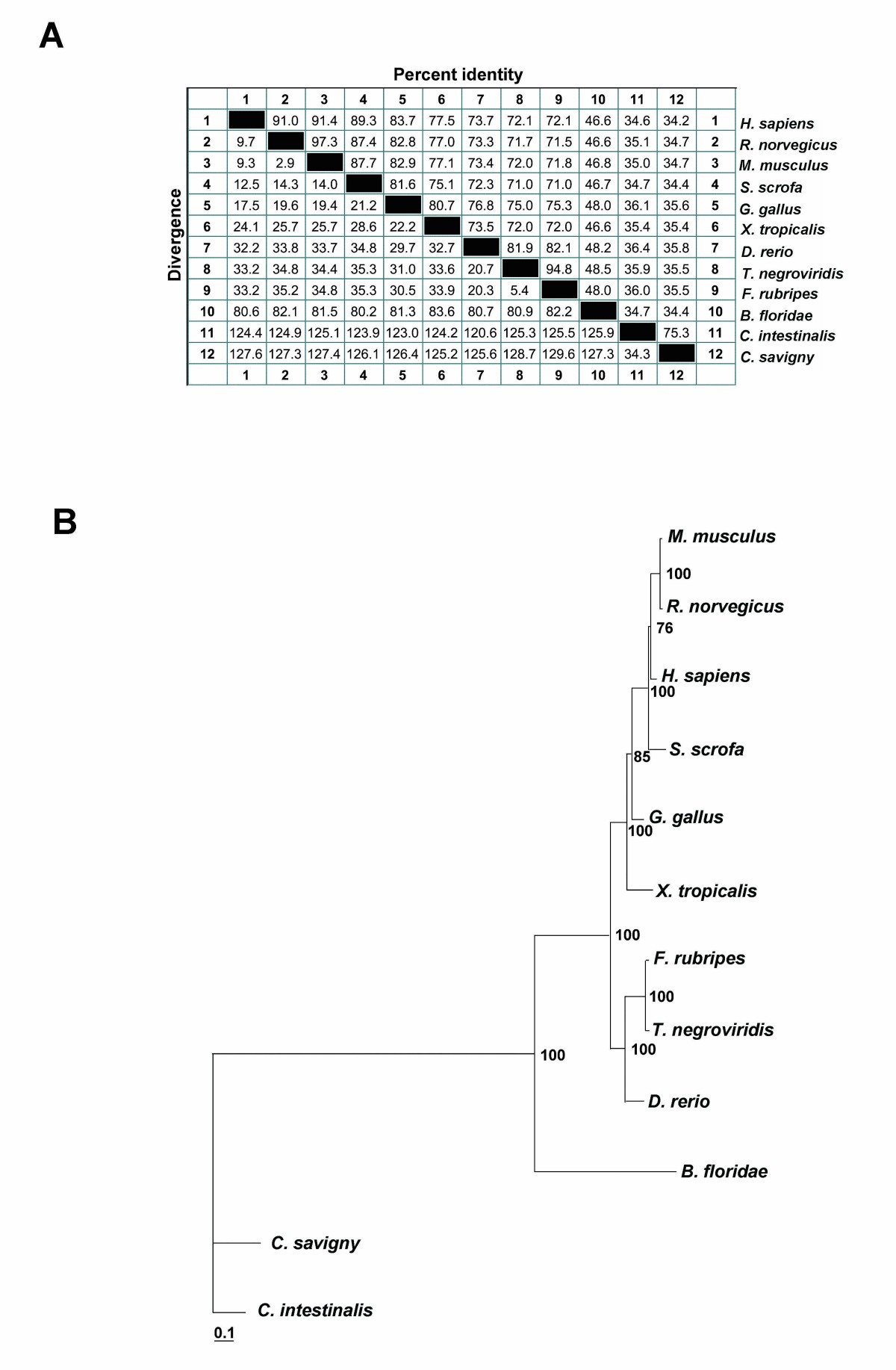
Characterization, developmental expression and evolutionary features of the huntingtin gene in the amphioxus Branchiostoma floridae | BMC Developmental Biology | Full Text
![PDF] TopModel: Template-based protein structure prediction at low sequence identity using top-down consensus and deep neural networks. | Semantic Scholar PDF] TopModel: Template-based protein structure prediction at low sequence identity using top-down consensus and deep neural networks. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/f7c338ed8a1fb5803453a17a953e2350c501ce0b/2-Figure1-1.png)
PDF] TopModel: Template-based protein structure prediction at low sequence identity using top-down consensus and deep neural networks. | Semantic Scholar

Protein sequence identity and similarity of BnCOMT1s. Data on the upper... | Download Scientific Diagram

How to find protein identity and similarities | Sequence homology | SIAS | Gene identity similarity - YouTube

Sequence-, structure-, and dynamics-based comparisons of structurally homologous CheY-like proteins | PNAS

Prediction of Protein Pairs Sharing Common Active Ligands Using Protein Sequence, Structure, and Ligand Similarity | Journal of Chemical Information and Modeling

NMR structures of two designed proteins with high sequence identity but different fold and function | PNAS

Difference Between Similarity and Identity in Sequence Alignment | Compare the Difference Between Similar Terms

AlphaFold 2 is here: what's behind the structure prediction miracle | Oxford Protein Informatics Group

Difference Between Similarity and Identity in Sequence Alignment | Compare the Difference Between Similar Terms

![PDF] Assessment of protein sequence identity from amino acid composition data. | Semantic Scholar PDF] Assessment of protein sequence identity from amino acid composition data. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/fd97264e93d2eb3dce2340fed985f4fb0b85eab1/6-Table1-1.png)