Measuring Illumina Size Bias Using REcount: A Novel Method for Highly Accurate Quantification of Engineered Genetic Constructs | bioRxiv
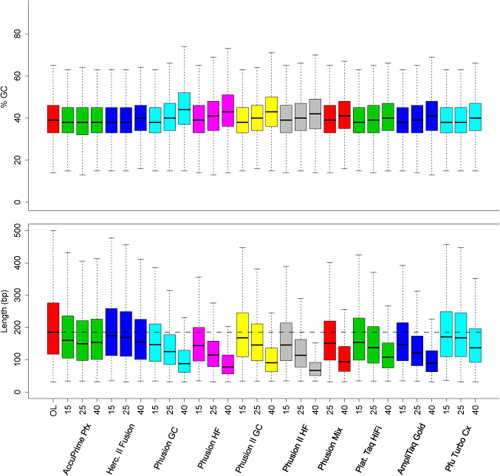
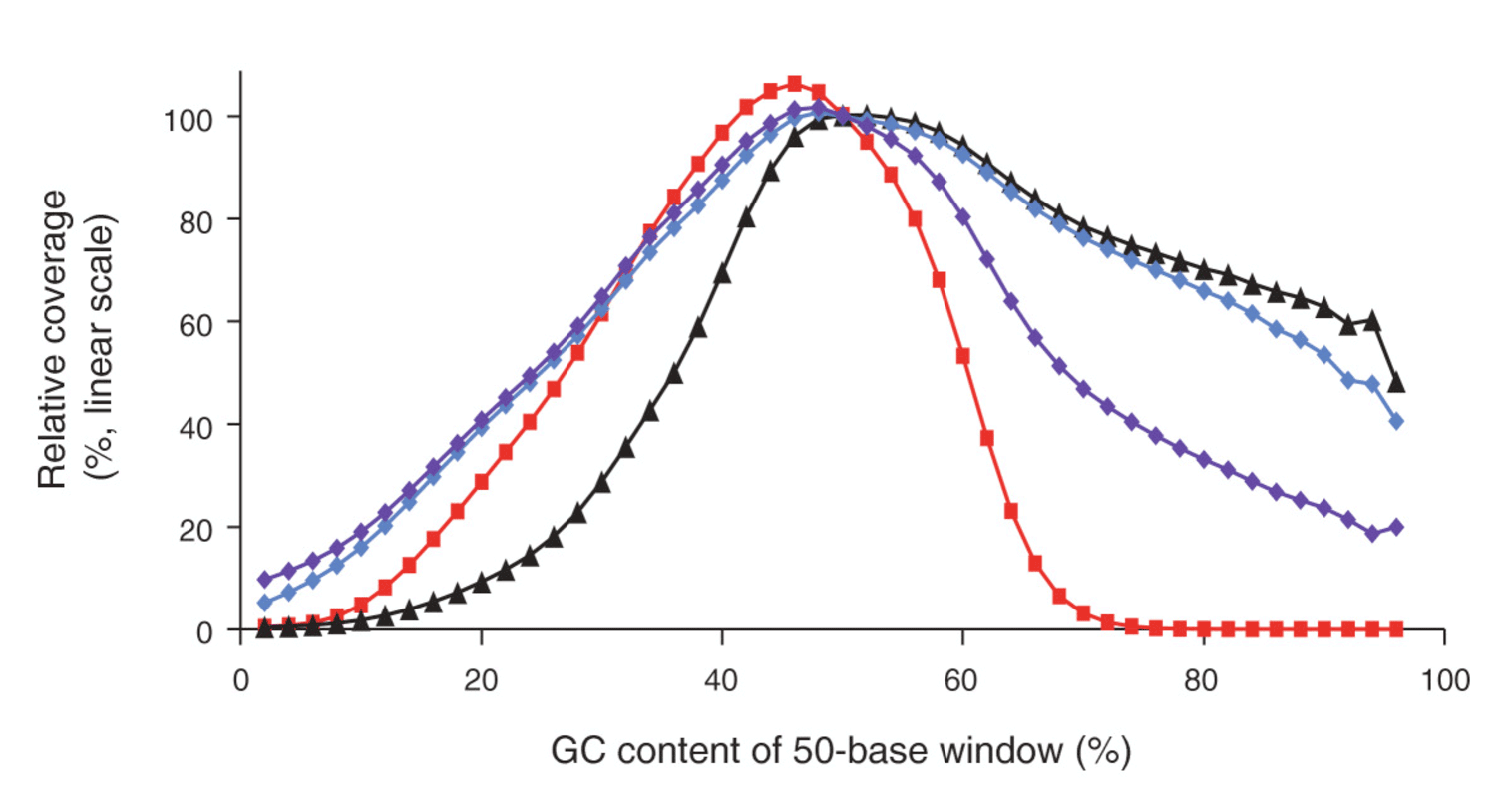
Length and GC-biases during sequencing library amplification: A comparison of various polymerase-buffer systems with ancient and modern DNA sequencing libraries | BioTechniques

Reply to Li et al., “GC Content-Associated Sequencing Bias Caused by Library Preparation Method May Infrequently Affect Salmonella Serotype Prediction Using SeqSero2” | Applied and Environmental Microbiology
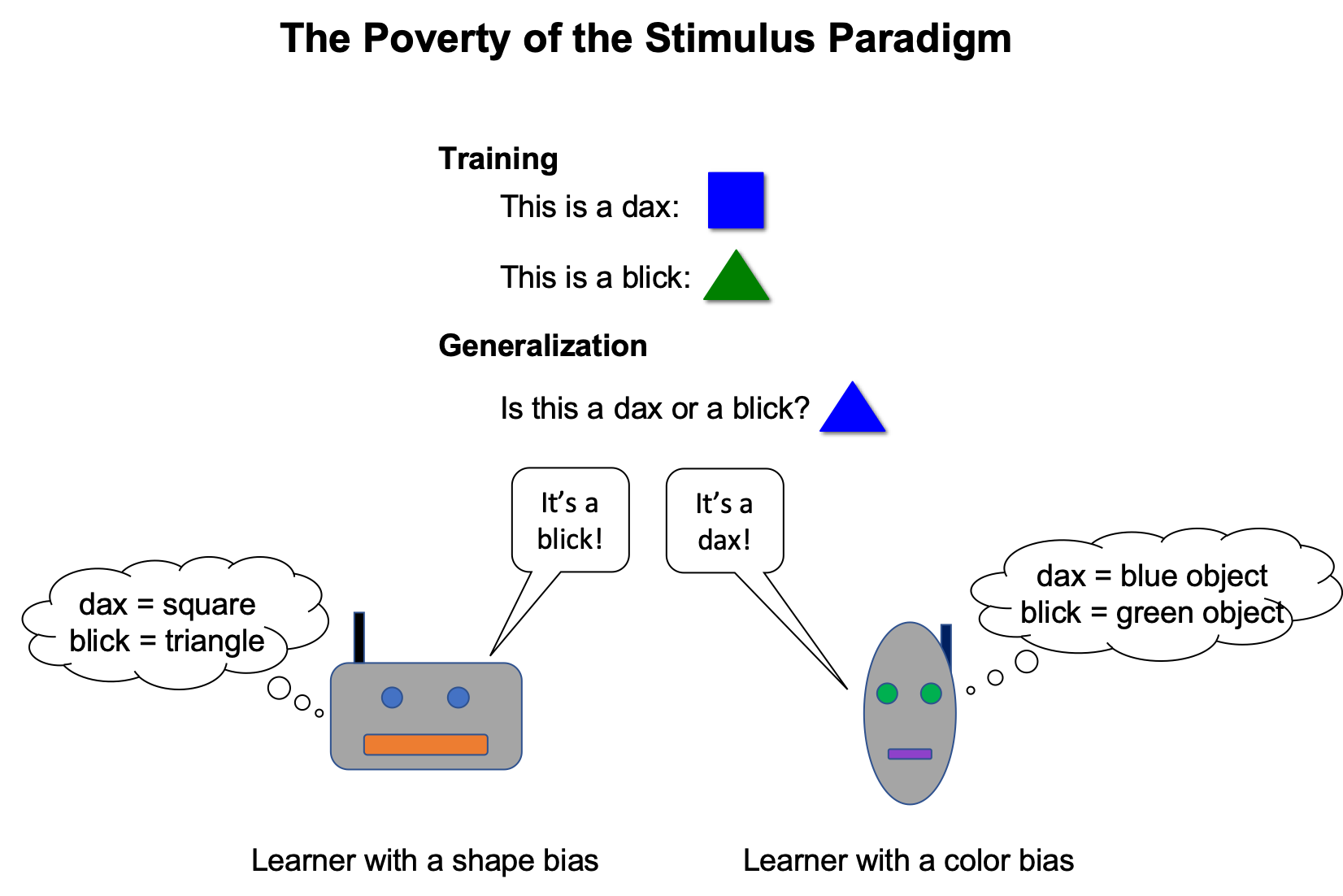
Does Syntax Need to Grow on Trees? Sources of Hierarchical Inductive Bias in Sequence-to-Sequence Networks

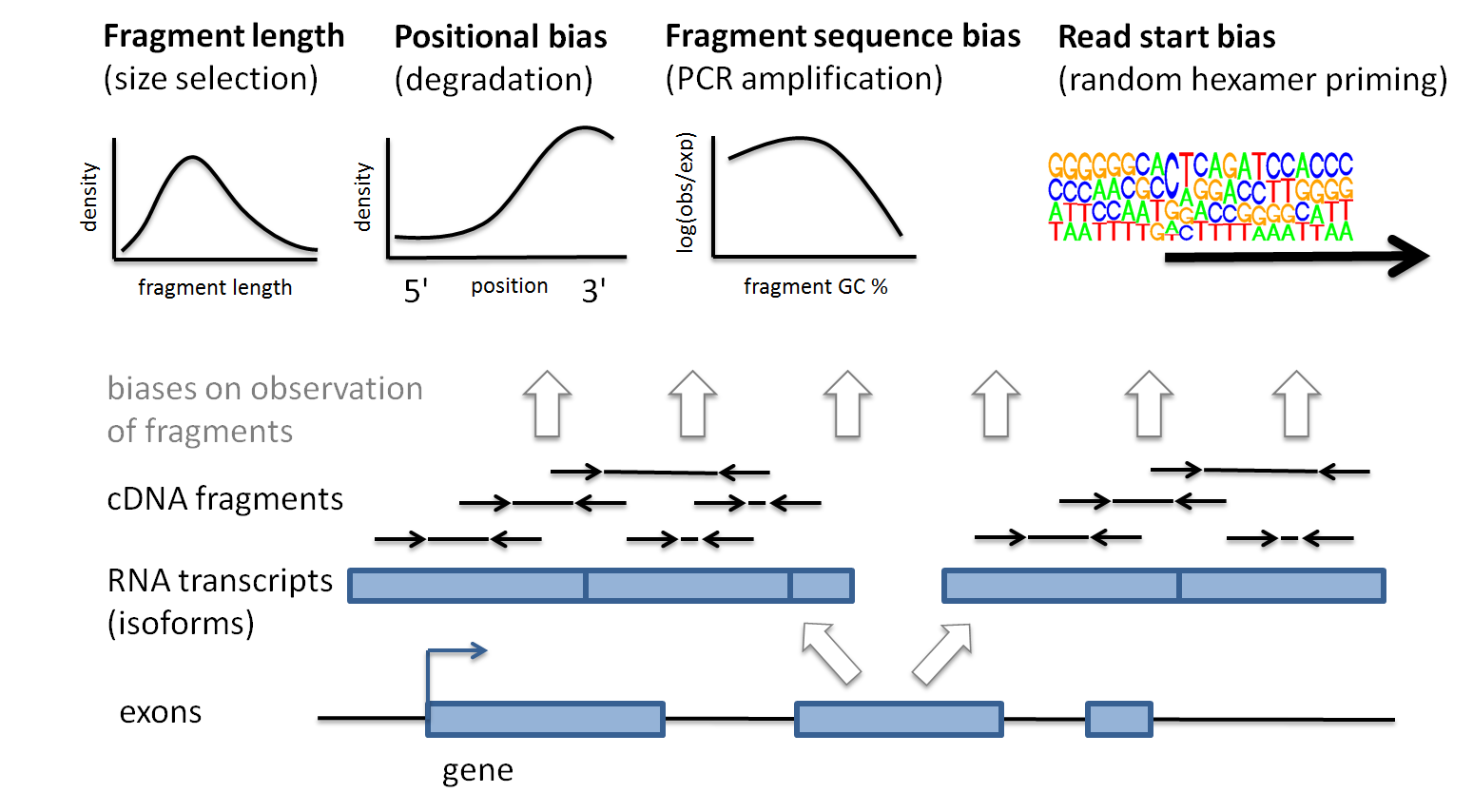
Modeling of RNA-seq fragment sequence bias reduces systematic errors in transcript abundance estimation | Nature Biotechnology
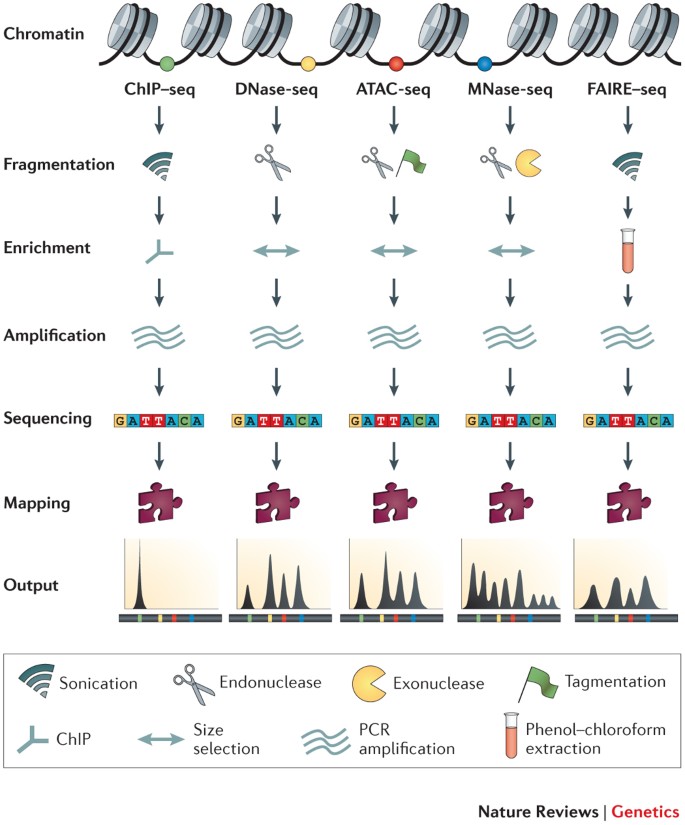
Identifying and mitigating bias in next-generation sequencing methods for chromatin biology | Nature Reviews Genetics

Sensitive and Low-Bias Transcriptome Sequencing Using Agarose PCR | ACS Applied Materials & Interfaces

Modeling Enzyme Processivity Reveals that RNA-Seq Libraries Are Biased in Characteristic and Correctable Ways: Cell Systems

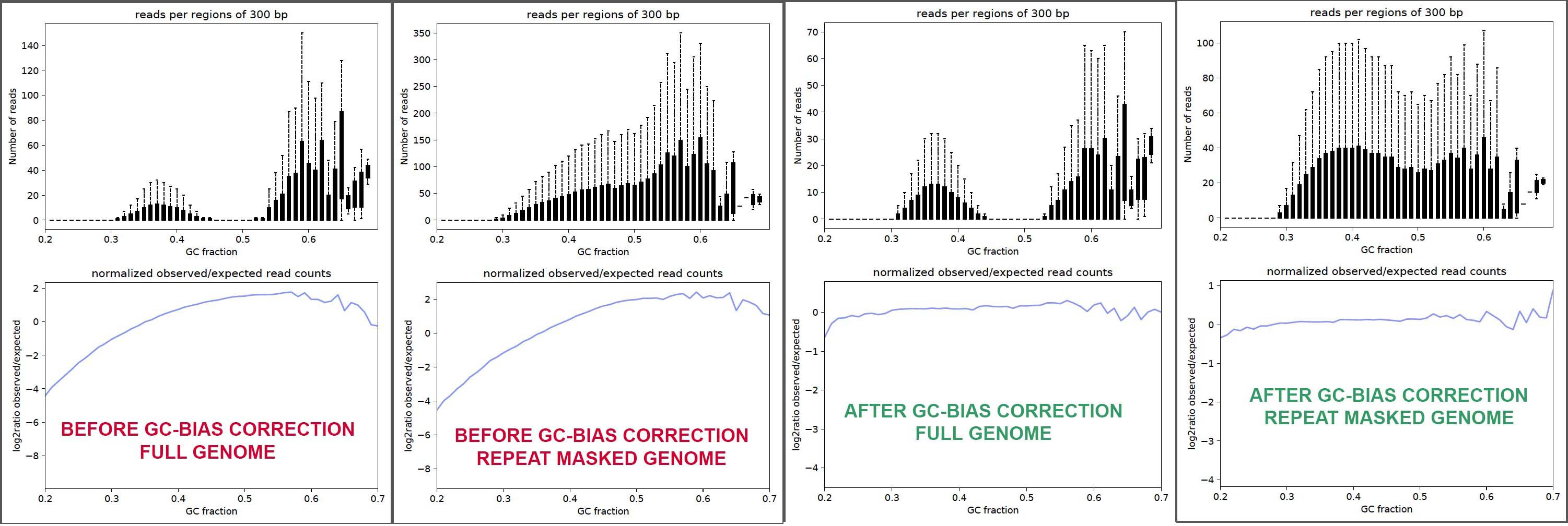
Figure 2 from Summarizing and correcting the GC content bias in high-throughput sequencing | Semantic Scholar