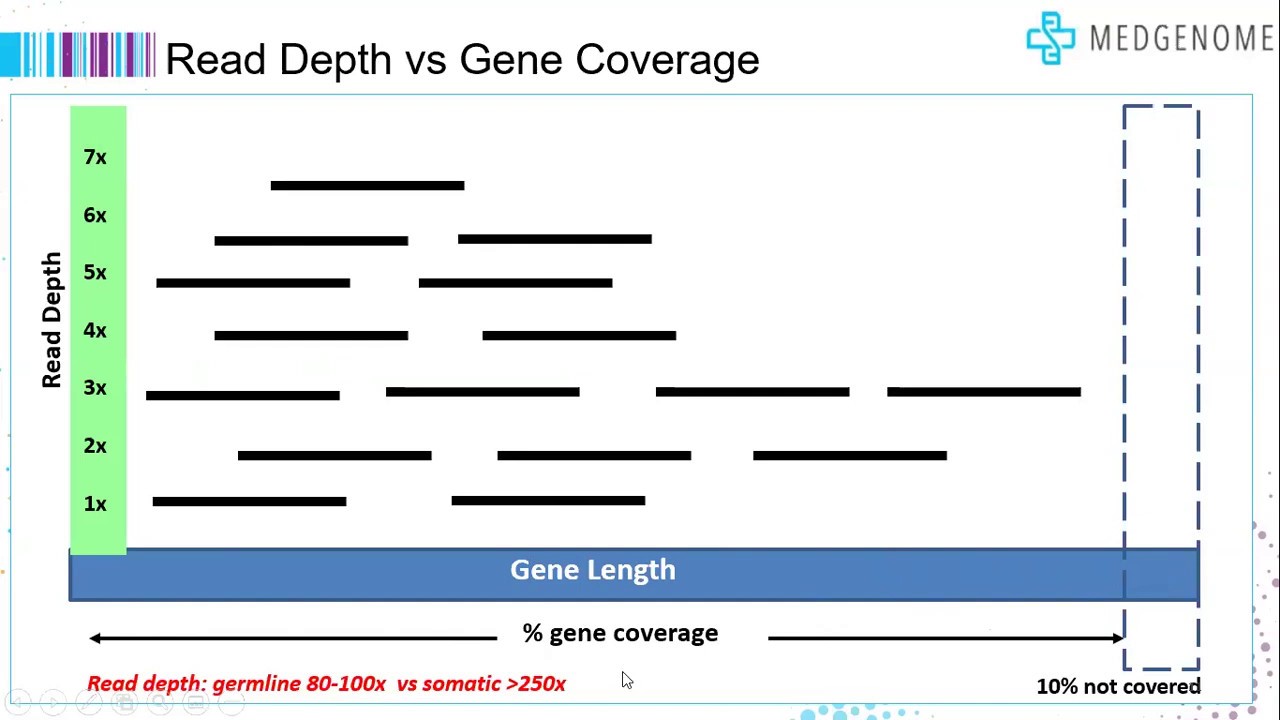
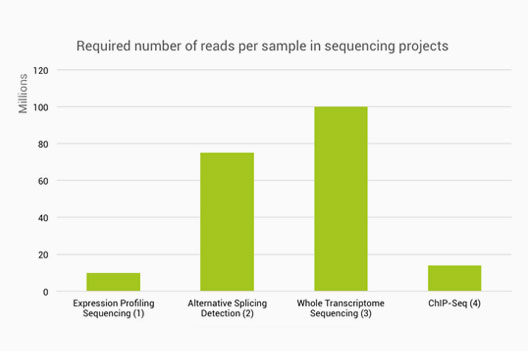
Standardization of Sequencing Coverage Depth in NGS: Recommendation for Detection of Clonal and Subclonal Mutations in Cancer Di
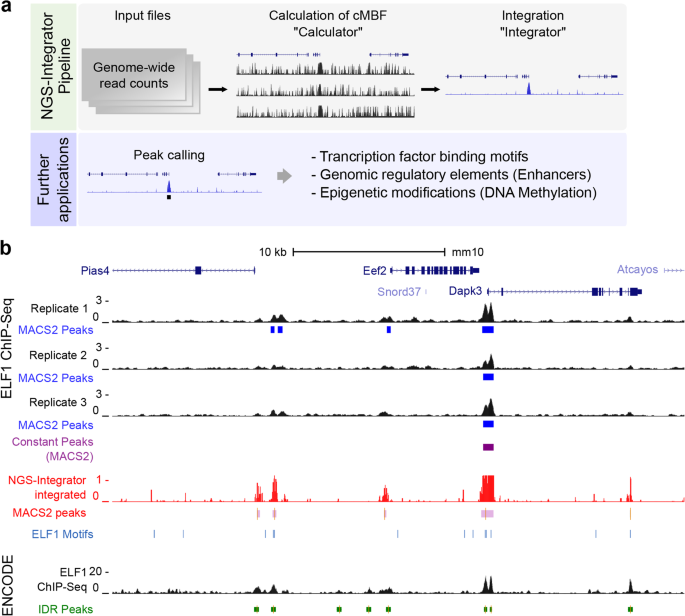
NGS-Integrator: An efficient tool for combining multiple NGS data tracks using minimum Bayes' factors | BMC Genomics | Full Text

Maximizing Sequence Coverage in Top-Down Proteomics By Automated Multimodal Gas-Phase Protein Fragmentation | Analytical Chemistry
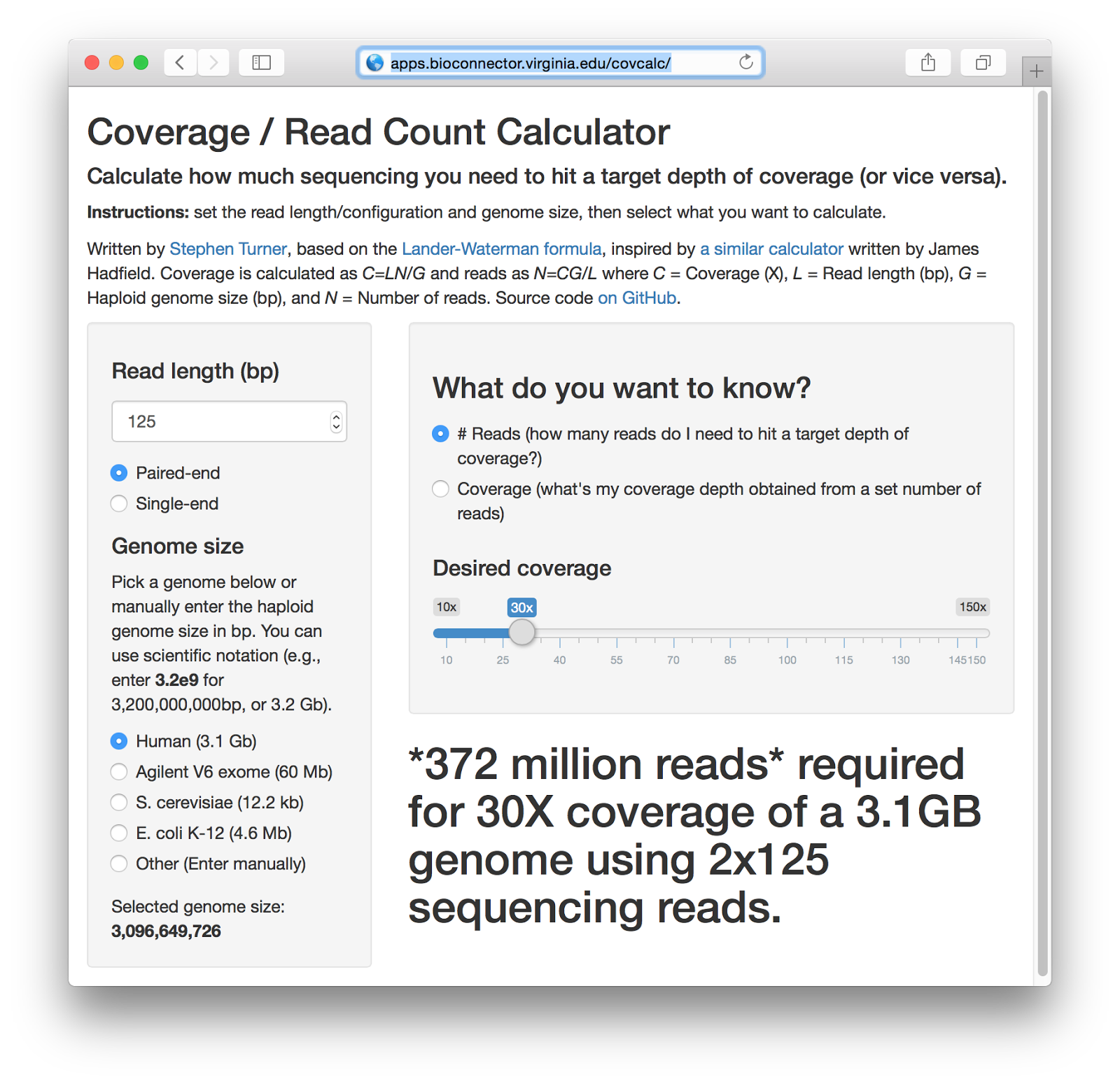
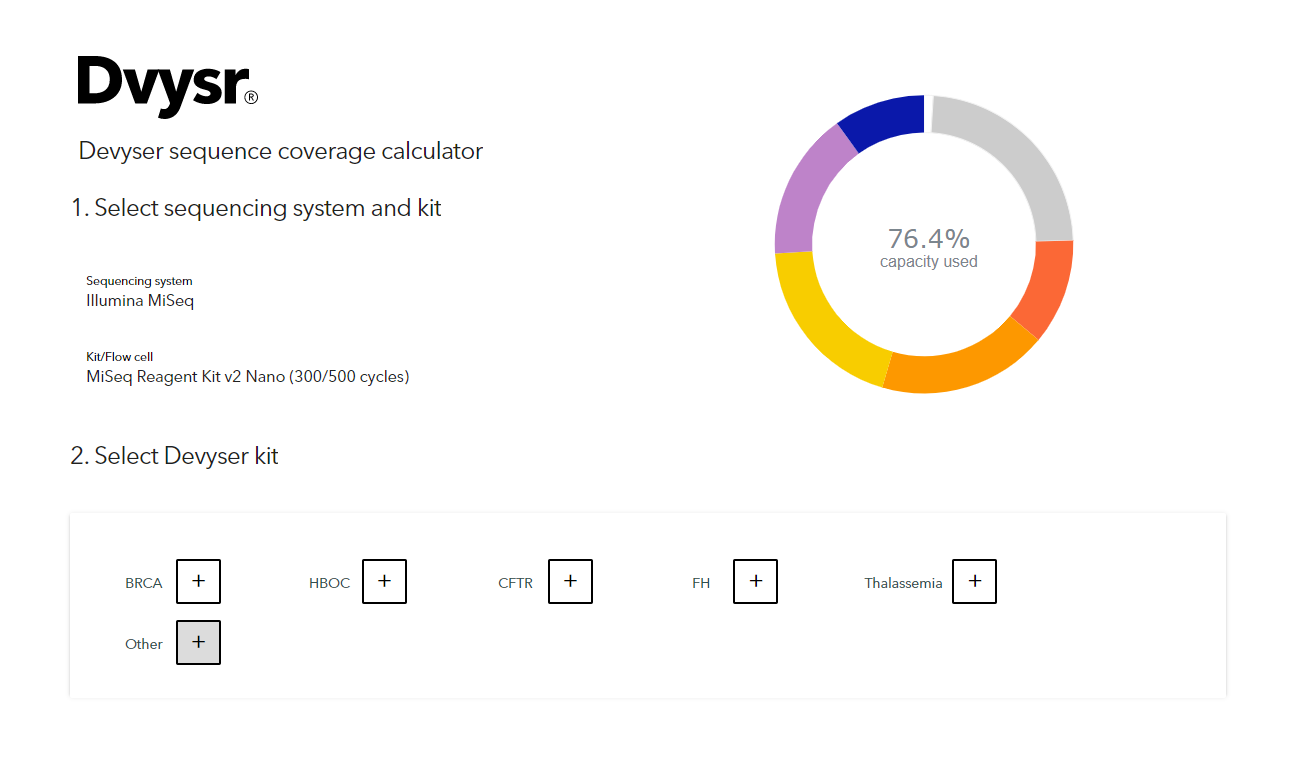
Getting Genetics Done: Covcalc: Shiny App for Calculating Coverage Depth or Read Counts for Sequencing Experiments

Mathematical Framework Provides a Read Depth Calculator and Guidelines... | Download Scientific Diagram
CNVkit: Genome-Wide Copy Number Detection and Visualization from Targeted DNA Sequencing | PLOS Computational Biology
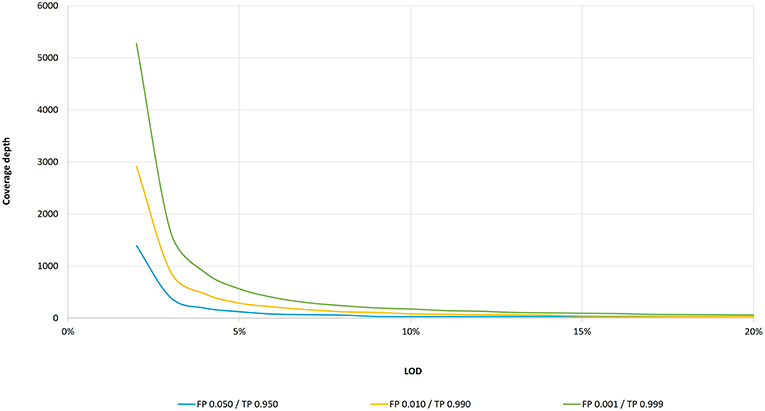
Frontiers | Standardization of Sequencing Coverage Depth in NGS: Recommendation for Detection of Clonal and Subclonal Mutations in Cancer Diagnostics
GitHub - GenomicaMicrob/coverage_calculator: A simple script to calculate the coverage of a genome assembly