![pGL4.73[hRluc /SV40] plasmid,pGL4.73[hRluc /SV40],pGL4.73[hRluc /SV40] plasmid,pGL4.73[hRluc /SV40] sequence,pGL4.73[hRluc /SV40] map pGL4.73[hRluc /SV40] plasmid,pGL4.73[hRluc /SV40],pGL4.73[hRluc /SV40] plasmid,pGL4.73[hRluc /SV40] sequence,pGL4.73[hRluc /SV40] map](https://lifescience-market.com/images/pGL4.73[hRluc-SV40].png)
pGL4.73[hRluc /SV40] plasmid,pGL4.73[hRluc /SV40],pGL4.73[hRluc /SV40] plasmid,pGL4.73[hRluc /SV40] sequence,pGL4.73[hRluc /SV40] map
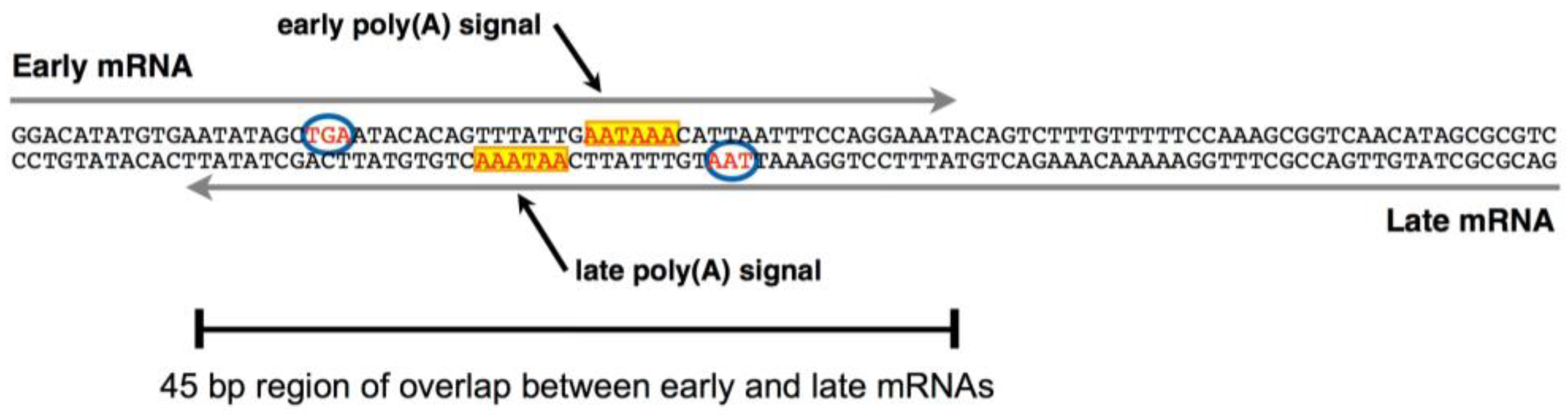
Sequences 5' to the polyadenylation signal mediate differential poly(A) site use in hepatitis B viruses

PiggyBac transposon-based polyadenylation-signal trap for genome-wide mutagenesis in mice | Scientific Reports

Assembly of the cleavage and polyadenylation apparatus requires about 10 seconds in vivo and is faster for strong than for weak poly(A) sites. - Abstract - Europe PMC

The Poly(A) Signal, without the Assistance of Any Downstream Element, Directs RNA Polymerase II to Pause in Vivo and Then to Release Stochastically from the Template* - Journal of Biological Chemistry

Comparison of polyadenylation signal sequences at the tailbud stage by... | Download Scientific Diagram

Schematic representation and nucleotide sequence of prothrombin variants expressed using the vectors harbouring the mutated SV40 polyadenylation signal
A functional .mRNA polyadeny.lation signal is required for transcnpnon termination by RNA polymerase II
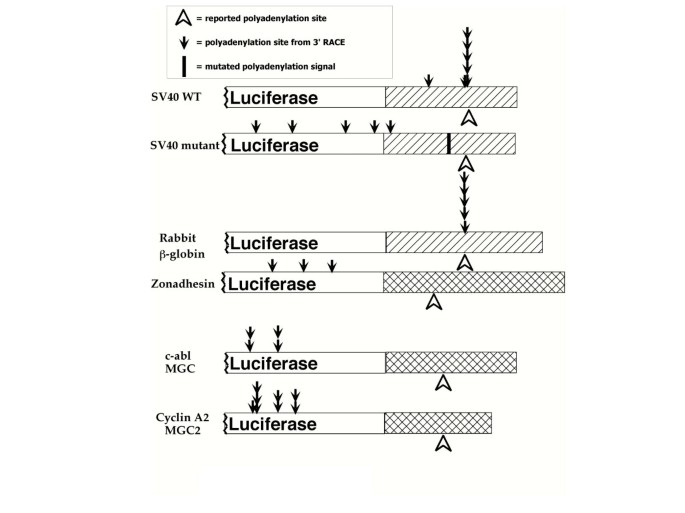
Differences in polyadenylation site choice between somatic and male germ cells | BMC Molecular Biology | Full Text