
Accurate Estimation of Molecular Counts from UMI-tagged Sequence data | Laurence H. Baker Center for Bioinformatics & Biological Statistics
Enabling high-accuracy long-read amplicon sequences using unique molecular identifiers and Nanopore sequencing
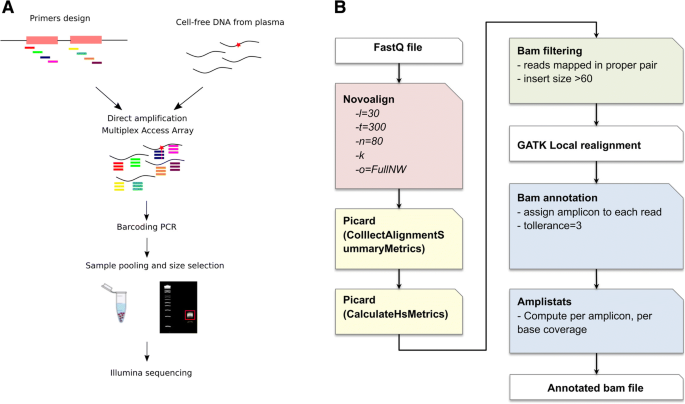
Next Generation-Targeted Amplicon Sequencing (NG-TAS): an optimised protocol and computational pipeline for cost-effective profiling of circulating tumour DNA | Genome Medicine | Full Text
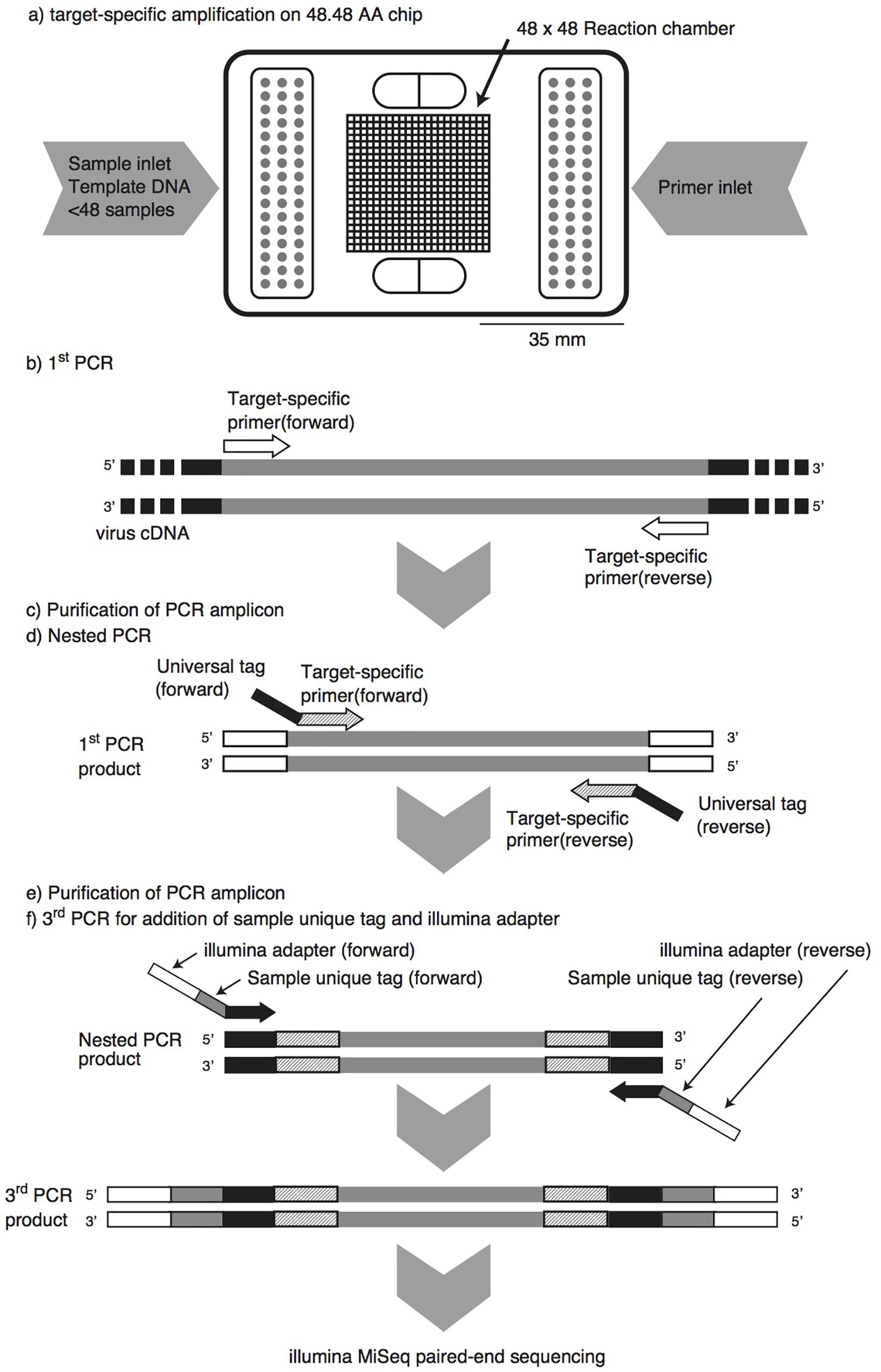
Frontiers | Microfluidic PCR Amplification and MiSeq Amplicon Sequencing Techniques for High-Throughput Detection and Genotyping of Human Pathogenic RNA Viruses in Human Feces, Sewage, and Oysters
of applications of Illumina amplicon sequencing for DNA barcoding and... | Download Scientific Diagram

Parallel tagged amplicon sequencing of relatively long PCR products using the Illumina HiSeq platform and transcriptome assembly - Feng - 2016 - Molecular Ecology Resources - Wiley Online Library

High-accuracy long-read amplicon sequences using unique molecular identifiers with Nanopore or PacBio sequencing | Nature Methods

Parallel tagged amplicon sequencing of relatively long PCR products using the Illumina HiSeq platform and transcriptome assembly - Feng - 2016 - Molecular Ecology Resources - Wiley Online Library
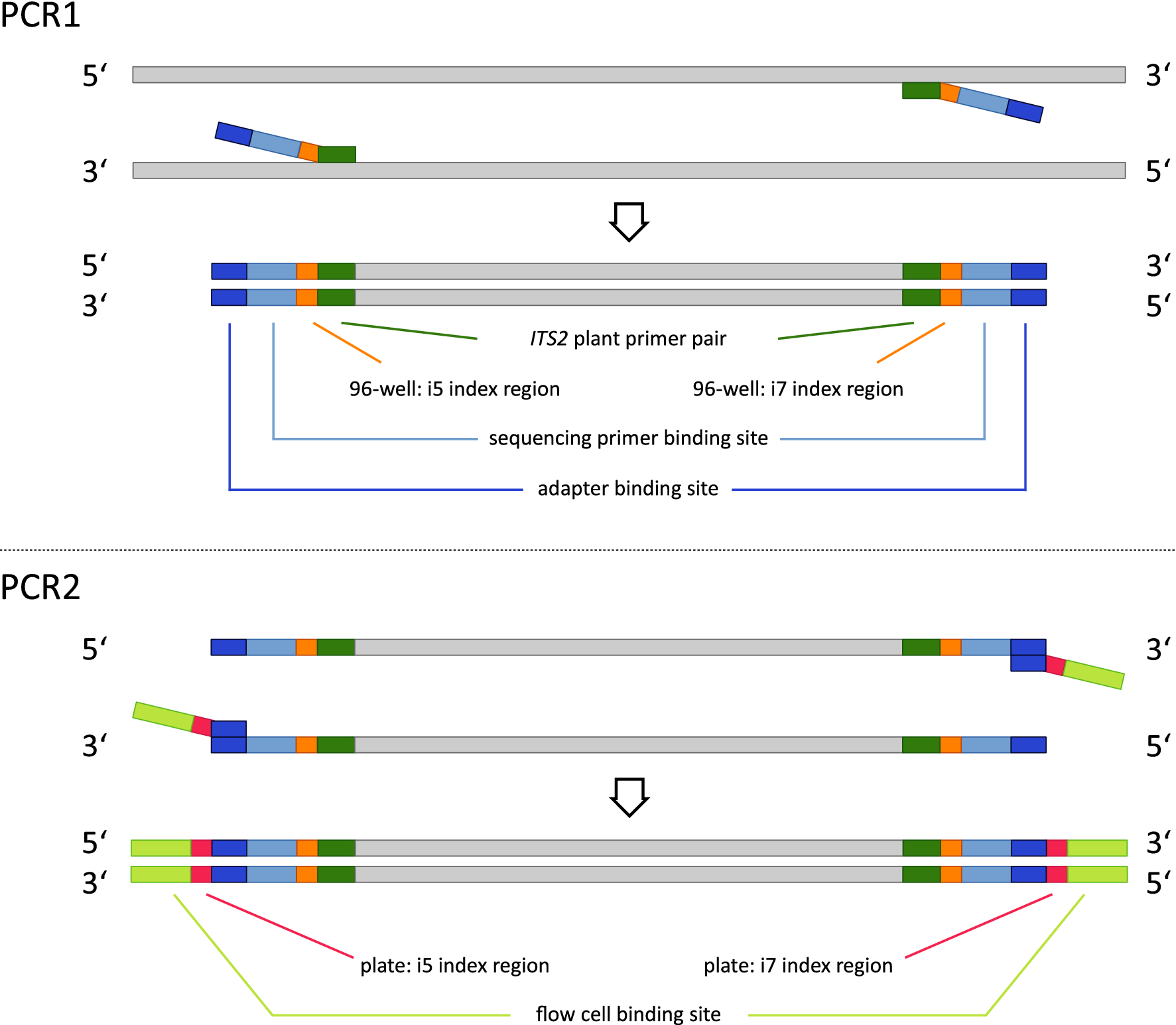
Handling of targeted amplicon sequencing data focusing on index hopping and demultiplexing using a nested metabarcoding approach in ecology | Scientific Reports
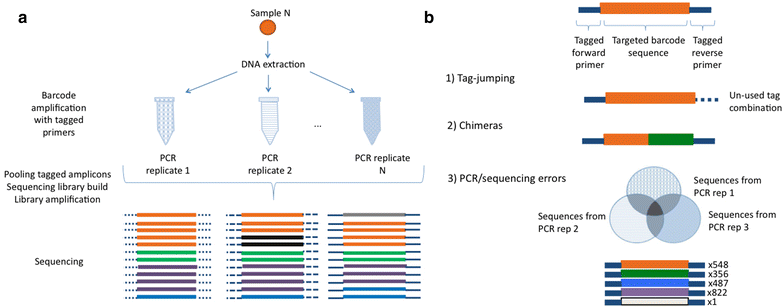
DAMe: a toolkit for the initial processing of datasets with PCR replicates of double-tagged amplicons for DNA metabarcoding analyses | BMC Research Notes | Full Text

Enabling high-accuracy long-read amplicon sequences using unique molecular identifiers with Nanopore or PacBio sequencing | bioRxiv

Multiplex amplicon sequencing for microbe identification in community-based culture collections | Scientific Reports
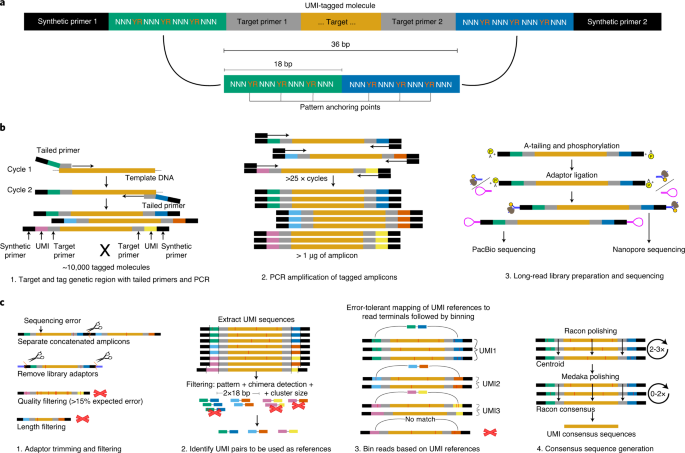
High-accuracy long-read amplicon sequences using unique molecular identifiers with Nanopore or PacBio sequencing | Nature Methods


![Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ] Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7786/1/fig-1-full.png)
![Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ] Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7786/1/fig-3-full.png)